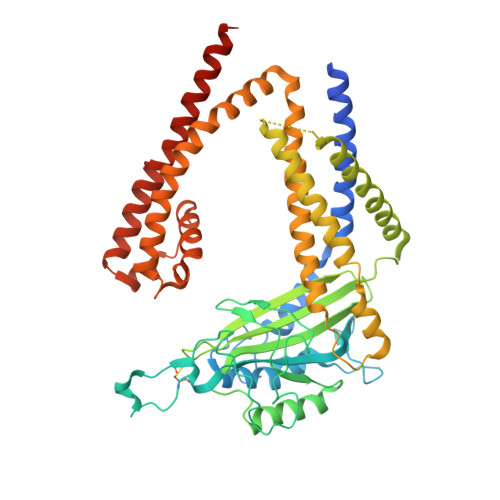
The Structure of the Polycystic Kidney Disease Channel PKD2 in Lipid Nanodiscs.
Shen, P.S., Yang, X., DeCaen, P.G., Liu, X., Bulkley, D., Clapham, D.E., Cao, E.(2016) Cell 167: 763-773.e11
- PubMed: 27768895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2016.09.048
- Primary Citation Related Structures:
5T4D - PubMed Abstract:
The Polycystic Kidney Disease 2 (Pkd2) gene is mutated in autosomal dominant polycystic kidney disease (ADPKD), one of the most common human monogenic disorders. Here, we present the cryo-EM structure of PKD2 in lipid bilayers at 3.0 Å resolution, which establishes PKD2 as a homotetrameric ion channel and provides insight into potential mechanisms for its activation. The PKD2 voltage-sensor domain retains two of four gating charges commonly found in those of voltage-gated ion channels. The PKD2 ion permeation pathway is constricted at the selectivity filter and near the cytoplasmic end of S6, suggesting that two gates regulate ion conduction. The extracellular domain of PKD2, a hotspot for ADPKD pathogenic mutations, contributes to channel assembly and strategically interacts with the transmembrane core, likely serving as a physical substrate for extracellular stimuli to allosterically gate the channel. Finally, our structure establishes the molecular basis for the majority of pathogenic mutations in Pkd2-related ADPKD.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, UT 84112-5650, USA.
Organizational Affiliation: