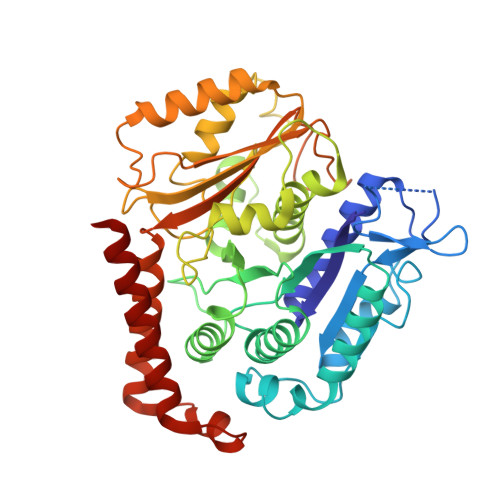
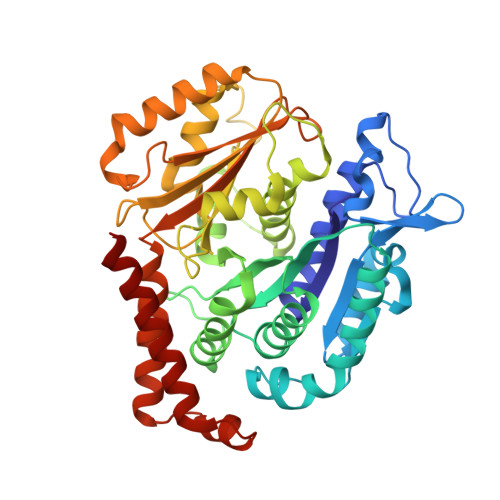
Insights into the Distinct Mechanisms of Action of Taxane and Non-Taxane Microtubule Stabilizers from Cryo-EM Structures.
Kellogg, E.H., Hejab, N.M., Howes, S., Northcote, P., Miller, J.H., Diaz, J.F., Downing, K.H., Nogales, E.(2017) J Mol Biology 429: 633-646
- PubMed: 28104363
- DOI: https://doi.org/10.1016/j.jmb.2017.01.001
- Primary Citation Related Structures:
5SYC, 5SYE, 5SYF, 5SYG - PubMed Abstract:
A number of microtubule (MT)-stabilizing agents (MSAs) have demonstrated or predicted potential as anticancer agents, but a detailed structural basis for their mechanism of action is still lacking. We have obtained high-resolution (3.9-4.2Å) cryo-electron microscopy (cryo-EM) reconstructions of MTs stabilized by the taxane-site binders Taxol and zampanolide, and by peloruside, which targets a distinct, non-taxoid pocket on β-tubulin. We find that each molecule has unique distinct structural effects on the MT lattice structure. Peloruside acts primarily at lateral contacts and has an effect on the "seam" of heterologous interactions, enforcing a conformation more similar to that of homologous (i.e., non-seam) contacts by which it regularizes the MT lattice. In contrast, binding of either Taxol or zampanolide induces MT heterogeneity. In doubly bound MTs, peloruside overrides the heterogeneity induced by Taxol binding. Our structural analysis illustrates distinct mechanisms of these drugs for stabilizing the MT lattice and is of relevance to the possible use of combinations of MSAs to regulate MT activity and improve therapeutic potential.
- Molecular Biophysics and Integrated Bioimaging Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.
Organizational Affiliation: