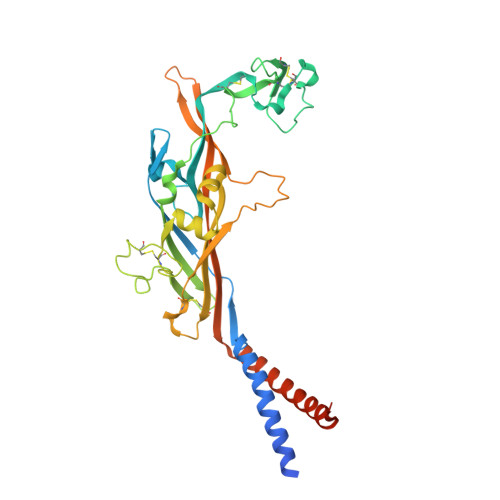
X-ray structures define human P2X3 receptor gating cycle and antagonist action.
Mansoor, S.E., Lu, W., Oosterheert, W., Shekhar, M., Tajkhorshid, E., Gouaux, E.(2016) Nature 538: 66-71
- PubMed: 27626375 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature19367
- Primary Citation Related Structures:
5SVJ, 5SVK, 5SVL, 5SVM, 5SVP, 5SVQ, 5SVR, 5SVS, 5SVT - PubMed Abstract:
P2X receptors are trimeric, non-selective cation channels activated by ATP that have important roles in the cardiovascular, neuronal and immune systems. Despite their central function in human physiology and although they are potential targets of therapeutic agents, there are no structures of human P2X receptors. The mechanisms of receptor desensitization and ion permeation, principles of antagonism, and complete structures of the pore-forming transmembrane domains of these receptors remain unclear. Here we report X-ray crystal structures of the human P2X 3 receptor in apo/resting, agonist-bound/open-pore, agonist-bound/closed-pore/desensitized and antagonist-bound/closed states. The open state structure harbours an intracellular motif we term the 'cytoplasmic cap', which stabilizes the open state of the ion channel pore and creates lateral, phospholipid-lined cytoplasmic fenestrations for water and ion egress. The competitive antagonists TNP-ATP and A-317491 stabilize the apo/resting state and reveal the interactions responsible for competitive inhibition. These structures illuminate the conformational rearrangements that underlie P2X receptor gating and provide a foundation for the development of new pharmacological agents.
- Vollum Institute, Oregon Health &Science University, Portland, Oregon 97239, USA.
Organizational Affiliation: