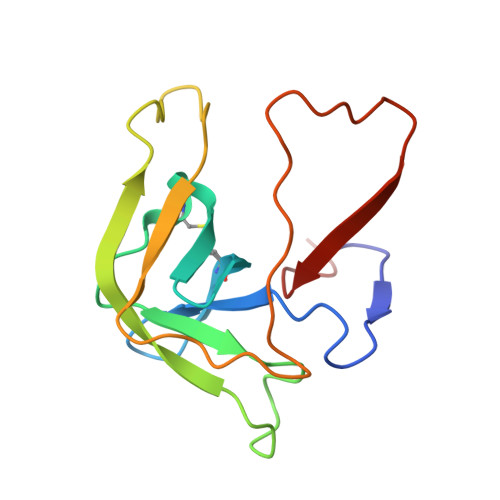
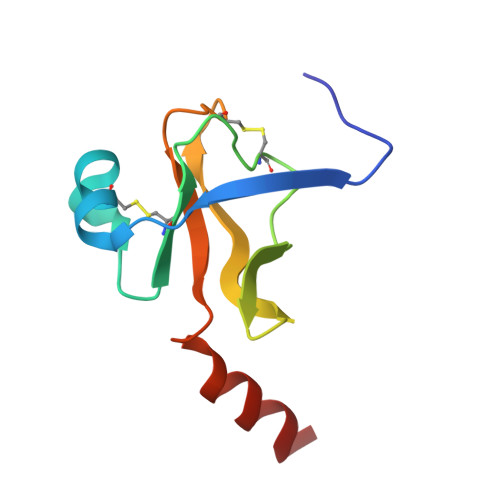
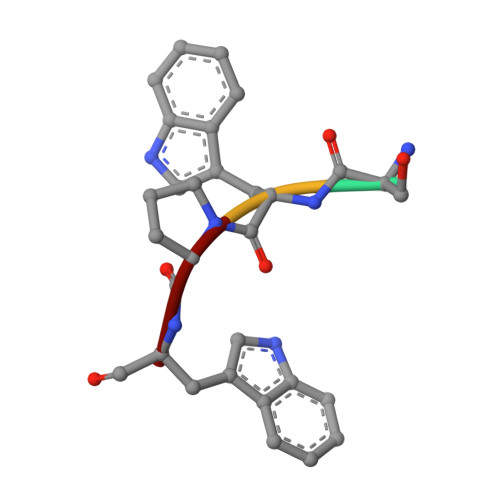
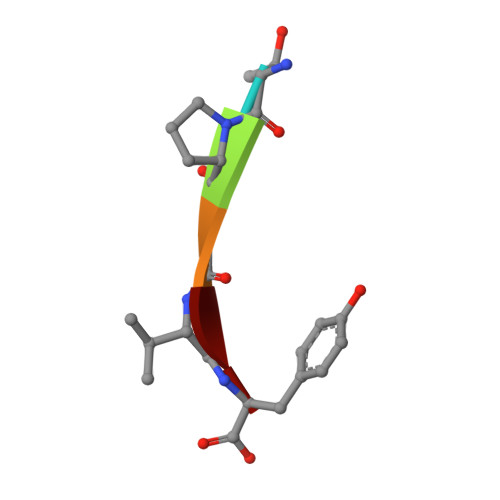
Effect of Temperature and pH on Ionizable Residues in gamma-Chymotrypsin: a X-ray and Neutron Crystallography Study
Kreinbring, C.A., Wilson, M.A., Kovalevsky, A.Y., Blakeley, M.P., Fisher, S.Z., Lazar, L.M., Moulin, A.G., Novak, W.R., Petsko, G.A., Ringe, D.To be published.