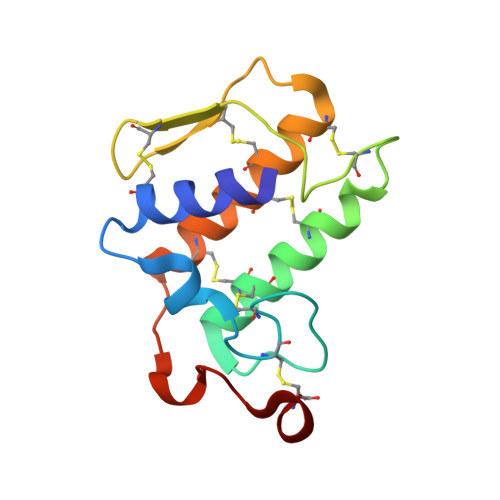
X-ray structure of phospholipase A2 complexed with a substrate-derived inhibitor.
Thunnissen, M.M., Ab, E., Kalk, K.H., Drenth, J., Dijkstra, B.W., Kuipers, O.P., Dijkman, R., de Haas, G.H., Verheij, H.M.(1990) Nature 347: 689-691
- PubMed: 2215698 Search on PubMed
- DOI: https://doi.org/10.1038/347689a0
- Primary Citation Related Structures:
5P2P - PubMed Abstract:
Phospholipases A2 play a part in a number of physiologically important cellular processes such as inflammation, blood platelet aggregation and acute hypersensitivity. These processes are all initiated by the release of arachidonic acid from cell membranes which is catalysed by intracellular phospholipases A2 and followed by conversion of arachidonic acid to prostaglandins, leukotrienes or thromboxanes. An imbalance in the production of these compounds can lead to chronic inflammatory diseases such as rheumatoid arthritis and asthma. Inhibitors of phospholipase A2 might therefore act to reduce the effects of inflammation, so structural information about the binding of phospholipase A2 to its substrates could be helpful in the design of therapeutic drugs. The three-dimensional structure is not known for any intracellular phospholipase A2, but these enzymes share significant sequence homology with secreted phospholipases, for which some of the structures have been determined. Here we report the structure of a complex between an extracellular phospholipase A2 and a competitively inhibiting substrate analogue, which reveals considerable detail about the interaction and suggests a mechanism for catalysis by this enzyme.
- Laboratory of Chemical Physics, University of Groningen, The Netherlands.
Organizational Affiliation: