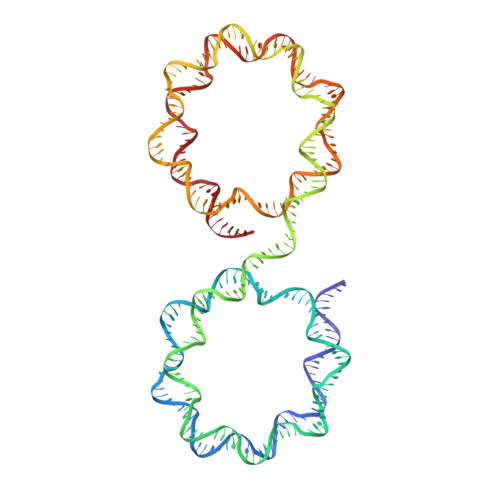
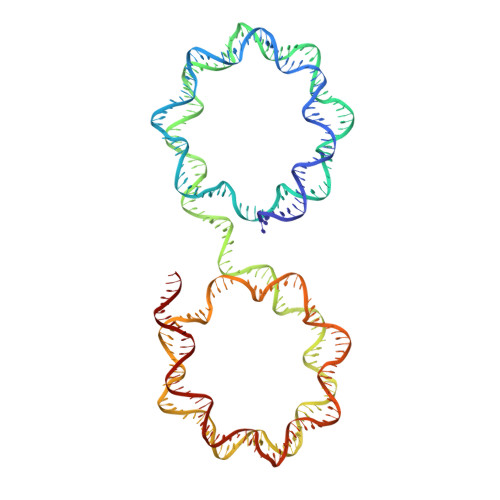
Capturing Structural Heterogeneity in Chromatin Fibers.
Ekundayo, B., Richmond, T.J., Schalch, T.(2017) J Mol Biology 429: 3031-3042
- PubMed: 28893533 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2017.09.002
- Primary Citation Related Structures:
5OXV, 5OY7 - PubMed Abstract:
Chromatin fiber organization is implicated in processes such as transcription, DNA repair and chromosome segregation, but how nucleosomes interact to form higher-order structure remains poorly understood. We solved two crystal structures of tetranucleosomes with approximately 11-bp DNA linker length at 5.8 and 6.7 Å resolution. Minimal intramolecular nucleosome-nucleosome interactions result in a fiber model resembling a flat ribbon that is compatible with a two-start helical architecture, and that exposes histone and DNA surfaces to the environment. The differences in the two structures combined with electron microscopy reveal heterogeneous structural states, and we used site-specific chemical crosslinking to assess the diversity of nucleosome-nucleosome interactions through identification of structure-sensitive crosslink sites that provide a means to characterize fibers in solution. The chromatin fiber architectures observed here provide a basis for understanding heterogeneous chromatin higher-order structures as they occur in a genomic context.
- Department of Molecular Biology, Faculty of Sciences, University of Geneva, CH-1211 Geneva 4, Switzerland; Institute of Genetics and Genomics of Geneva (iGE3), University of Geneva, CH-1211 Geneva 4, Switzerland.
Organizational Affiliation: