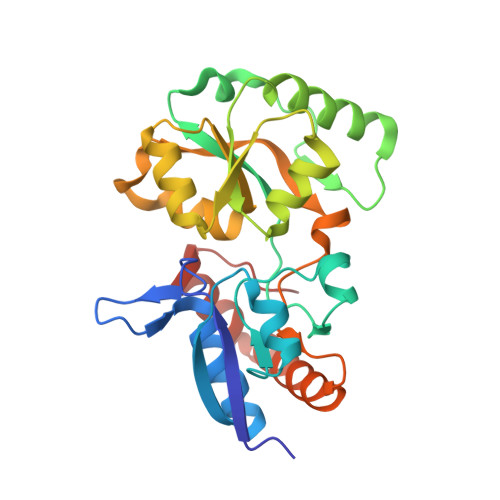
Structural basis for high specificity of octopine binding in the plant pathogen Agrobacterium tumefaciens.
Vigouroux, A., El Sahili, A., Lang, J., Aumont-Nicaise, M., Dessaux, Y., Faure, D., Morera, S.(2017) Sci Rep 7: 18033-18033
- PubMed: 29269740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-18243-8
- Primary Citation Related Structures:
5ORE, 5ORG, 5OT8, 5OT9, 5OTA, 5OTC - PubMed Abstract:
Agrobacterium pathogens of octopine- and nopaline-types force host plants to produce either octopine or nopaline compounds, which they use as nutrients. Two Agrobacterium ABC-transporters and their cognate periplasmic binding proteins (PBPs) OccJ and NocT import octopine and nopaline/octopine, respectively. Here, we show that both octopine transport and degradation confer a selective advantage to octopine-type A. tumefaciens when it colonizes plants. We report the X-ray structures of the unliganded PBP OccJ and its complex with octopine as well as a structural comparison with NocT and the related PBP LAO from Salmonella enterica, which binds amino acids (lysine, arginine and ornithine). We investigated the specificity of OccJ, NocT and LAO using several ligands such as amino acids, octopine, nopaline and octopine analogues. OccJ displays a high selectivity and nanomolar range affinity for octopine. Altogether, the structural and affinity data allowed to define an octopine binding signature in PBPs and to construct a OccJ mutant impaired in octopine binding, a selective octopine-binding NocT and a non-selective octopine-binding LAO by changing one single residue in these PBPs. We proposed the PBP OccJ as a major trait in the ecological specialization of octopine-type Agrobacterium pathogens when they colonize and exploit the plant host.
- Institute for Integrative Biology of the Cell (I2BC), CNRS CEA Univ. Paris-Sud, Université Paris-Saclay, Avenue de la Terrasse, Gif-sur-Yvette, 91198, France.
Organizational Affiliation: