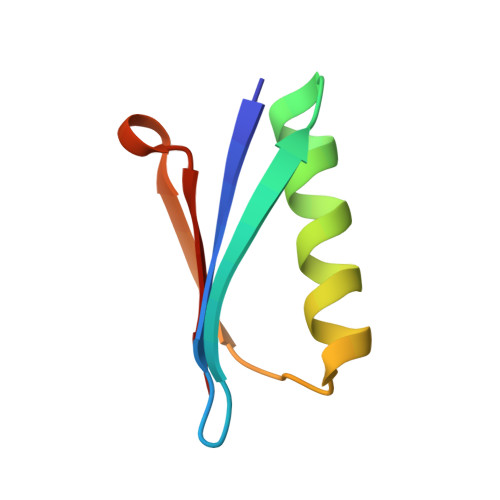
Genetic Algorithm Based Design and Experimental Characterization of a Highly Thermostable Metalloprotein.
Bozkurt, E., Perez, M.A.S., Hovius, R., Browning, N.J., Rothlisberger, U.(2018) J Am Chem Soc 140: 4517-4521
- PubMed: 29336153 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.7b10660
- Primary Citation Related Structures:
5O94, 5OFS - PubMed Abstract:
The development of thermostable and solvent-tolerant metalloproteins is a long-sought goal for many applications in synthetic biology and biotechnology. In this work, we were able to engineer a highly thermostable and organic solvent-stable metallo variant of the B1 domain of protein G (GB1) with a tetrahedral zinc binding site reminiscent of the one of thermolysin. Promising candidates were designed computationally by applying a protocol based on classical and first-principles molecular dynamics simulations in combination with genetic algorithm optimization. The most promising of the computationally predicted mutants was expressed and structurally characterized and yielded a highly thermostable protein. The experimental results thus confirm the predictive power of the applied computational protein engineering approach for the de novo design of highly stable metalloproteins.
- Laboratory of Computational Chemistry and Biochemistry , École Polytechnique Fédérale de Lausanne , CH-1015 Lausanne , Switzerland.
Organizational Affiliation: