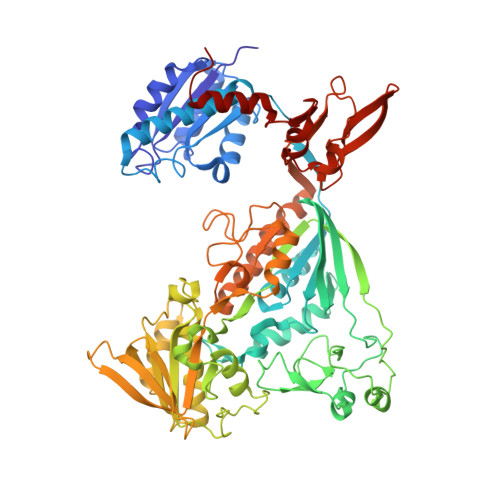
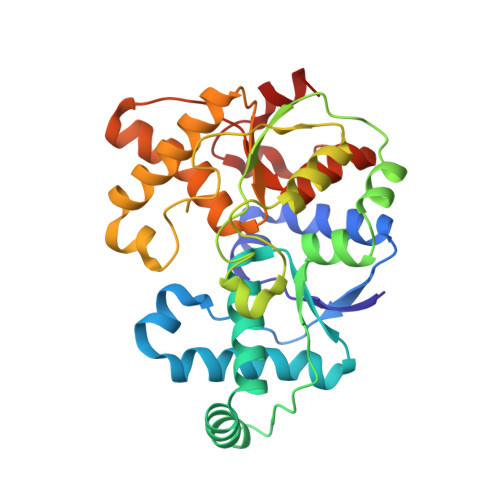
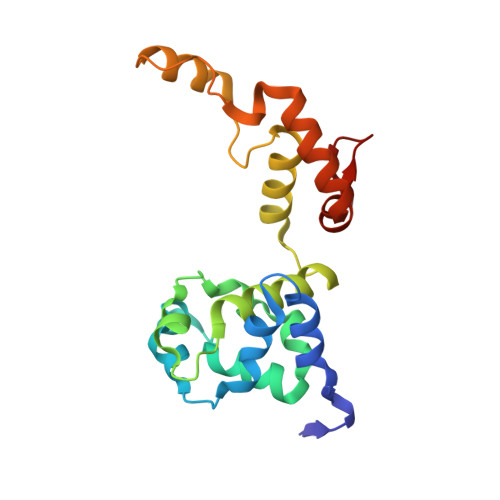
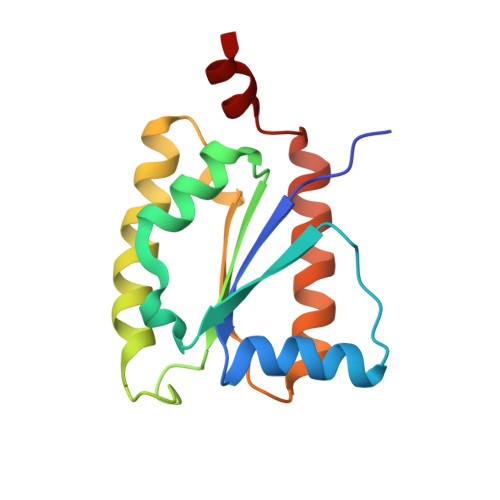
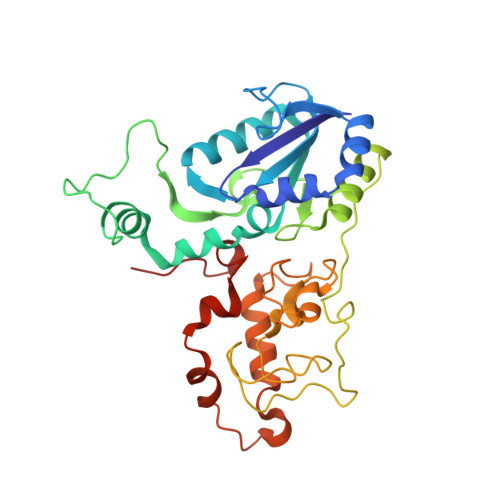
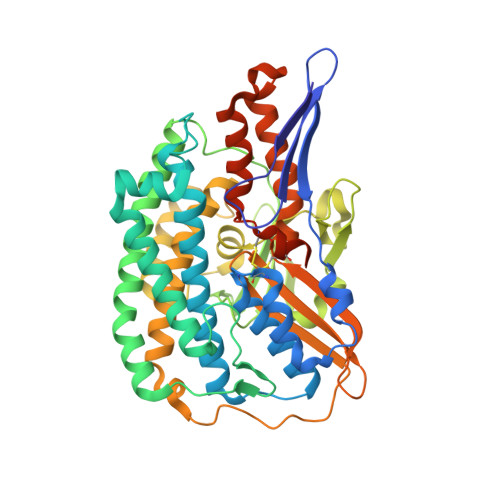
Methanogenic heterodisulfide reductase (HdrABC-MvhAGD) uses two noncubane [4Fe-4S] clusters for reduction.
Wagner, T., Koch, J., Ermler, U., Shima, S.(2017) Science 357: 699-703
- PubMed: 28818947 Search on PubMed
- DOI: https://doi.org/10.1126/science.aan0425
- Primary Citation Related Structures:
5ODC, 5ODH, 5ODI, 5ODQ, 5ODR - PubMed Abstract:
In methanogenic archaea, the carbon dioxide (CO 2 ) fixation and methane-forming steps are linked through the heterodisulfide reductase (HdrABC)-[NiFe]-hydrogenase (MvhAGD) complex that uses flavin-based electron bifurcation to reduce ferredoxin and the heterodisulfide of coenzymes M and B. Here, we present the structure of the native heterododecameric HdrABC-MvhAGD complex at 2.15-angstrom resolution. HdrB contains two noncubane [4Fe-4S] clusters composed of fused [3Fe-4S]-[2Fe-2S] units sharing 1 iron (Fe) and 1 sulfur (S), which were coordinated at the CCG motifs. Soaking experiments showed that the heterodisulfide is clamped between the two noncubane [4Fe-4S] clusters and homolytically cleaved, forming coenzyme M and B bound to each iron. Coenzymes are consecutively released upon one-by-one electron transfer. The HdrABC-MvhAGD atomic model serves as a structural template for numerous HdrABC homologs involved in diverse microbial metabolic pathways.
- Max Planck Institute for Terrestrial Microbiology, Karl-von-Frisch-Straße 10, 35043 Marburg, Germany.
Organizational Affiliation: