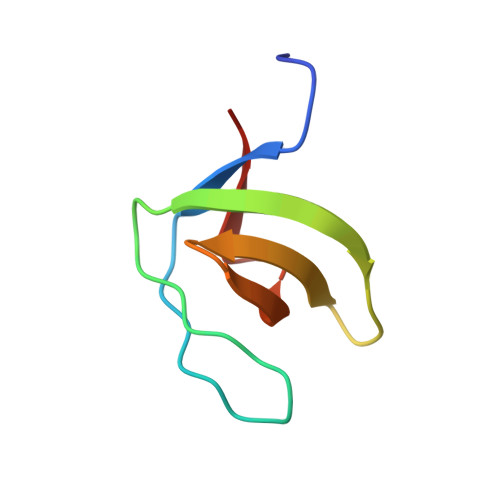
Structure of an unconventional SH3 domain from the postsynaptic density protein Shank3 at ultrahigh resolution.
Ponna, S.K., Myllykoski, M., Boeckers, T.M., Kursula, P.(2017) Biochem Biophys Res Commun 490: 806-812
- PubMed: 28647360 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.06.121
- Primary Citation Related Structures:
5O99 - PubMed Abstract:
The Shank family comprises three large multi-domain proteins playing central roles as protein scaffolds in the neuronal postsynaptic density. The Shank proteins are closely linked to neuropsychiatric diseases, such as autism spectrum disorders. One characteristic domain in the Shank family is the SH3 domain, assumed to play a role in protein-protein interactions; however, no specific ligand binding to any Shank SH3 domain has been described. We solved the crystal structure of the SH3 domain from Shank3 at sub-atomic resolution. While the structure presents the canonical SH3 domain fold, the binding site for proline-rich peptides is not conserved. In line with this, no binding of Pro-rich sequences by the Shank3 SH3 domain was observed. Sequence comparisons indicate that all Shank isoforms have similarly lost the classical Pro-rich peptide binding site from the SH3 domain. Whether the corresponding site in the Shank SH3 domains has evolved to bind a non-poly-Pro target sequence is currently not known. Our work provides an intriguing example of the evolution of a well-characterized protein-protein interaction domain within the context of multi-domain protein scaffolds, allowing the conservation of structural features, but losing canonical functional sites. The data are further discussed in light of known mutations in the SH3 domain or its vicinity in the different Shank isoforms.
- Faculty of Biochemistry and Molecular Medicine & Biocenter Oulu, University of Oulu, Finland.
Organizational Affiliation: