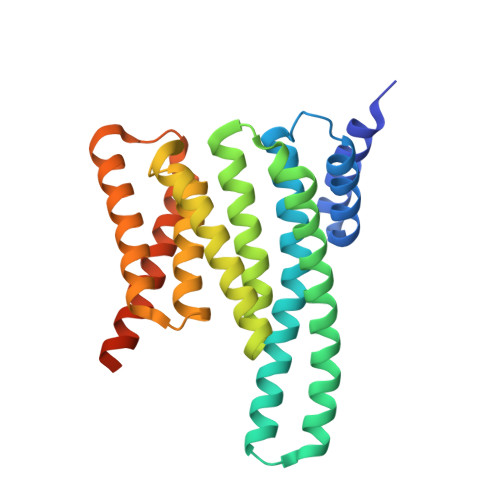
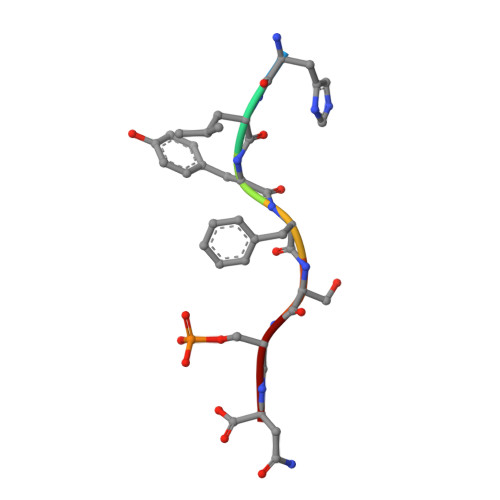
Fusicoccin Activates KAT1 Channels by Stabilizing Their Interaction with 14-3-3 Proteins.
Saponaro, A., Porro, A., Chaves-Sanjuan, A., Nardini, M., Rauh, O., Thiel, G., Moroni, A.(2017) Plant Cell 29: 2570-2580
- PubMed: 28970335 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1105/tpc.17.00375
- Primary Citation Related Structures:
5NWI, 5NWJ, 5NWK - PubMed Abstract:
Plants acquire potassium (K + ) ions for cell growth and movement via regulated diffusion through K + channels. Here, we present crystallographic and functional data showing that the K + inward rectifier KAT1 (K + Arabidopsis thaliana 1) channel is regulated by 14-3-3 proteins and further modulated by the phytotoxin fusicoccin, in analogy to the H + -ATPase. We identified a 14-3-3 mode III binding site at the very C terminus of KAT1 and cocrystallized it with tobacco ( Nicotiana tabacum ) 14-3-3 proteins to describe the protein complex at atomic detail. Validation of this interaction by electrophysiology shows that 14-3-3 binding augments KAT1 conductance by increasing the maximal current and by positively shifting the voltage dependency of gating. Fusicoccin potentiates the 14-3-3 effect on KAT1 activity by stabilizing their interaction. Crystal structure of the ternary complex reveals a noncanonical binding site for the toxin that adopts a novel conformation. The structural insights underscore the adaptability of fusicoccin, predicting more potential targets than so far anticipated. The data further advocate a common mechanism of regulation of the proton pump and a potassium channel, two essential elements in K + uptake in plant cells.
- Department of Biosciences, University of Milan, 20133 Milan, Italy.
Organizational Affiliation: