Group-Based Optimization of Potent and Cell-Active Inhibitors of the von Hippel-Lindau (VHL) E3 Ubiquitin Ligase: Structure-Activity Relationships Leading to the Chemical Probe (2S,4R)-1-((S)-2-(1-Cyanocyclopropanecarboxamido)-3,3-dimethylbutanoyl)-4-hydroxy-N-(4-(4-methylthiazol-5-yl)benzyl)pyrrolidine-2-carboxamide (VH298).
Soares, P., Gadd, M.S., Frost, J., Galdeano, C., Ellis, L., Epemolu, O., Rocha, S., Read, K.D., Ciulli, A.(2018) J Med Chem 61: 599-618
- PubMed: 28853884 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.7b00675
- Primary Citation Related Structures:
5NVV, 5NVW, 5NVX, 5NVY, 5NVZ, 5NW0, 5NW1, 5NW2 - PubMed Abstract:
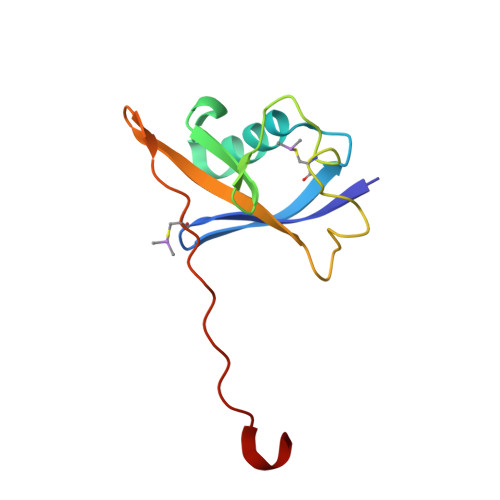
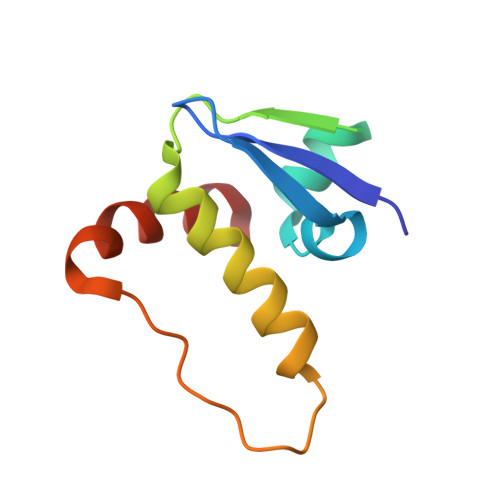
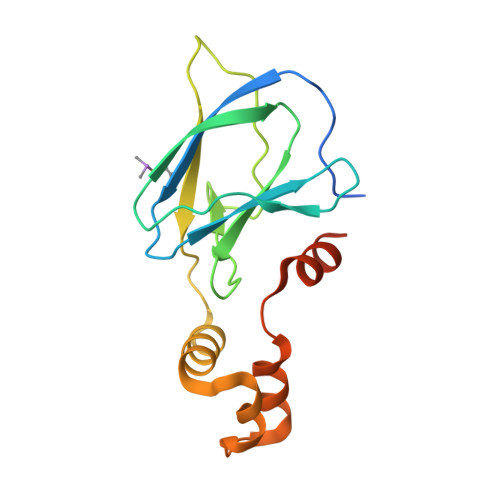
The von Hippel-Lindau tumor suppressor protein is the substrate binding subunit of the VHL E3 ubiquitin ligase, which targets hydroxylated α subunit of hypoxia inducible factors (HIFs) for ubiquitination and subsequent proteasomal degradation. VHL is a potential target for treating anemia and ischemic diseases, motivating the development of inhibitors of the VHL:HIF-α protein-protein interaction. Additionally, bifunctional proteolysis targeting chimeras (PROTACs) containing a VHL ligand can hijack the E3 ligase activity to induce degradation of target proteins. We report the structure-guided design and group-based optimization of a series of VHL inhibitors with low nanomolar potencies and improved cellular permeability. Structure-activity relationships led to the discovery of potent inhibitors 10 and chemical probe VH298, with dissociation constants <100 nM, which induced marked HIF-1α intracellular stabilization. Our study provides new chemical tools to probe the VHL-HIF pathways and new VHL ligands for next-generation PROTACs.
- Division of Biological Chemistry and Drug Discovery, School of Life Sciences, University of Dundee , Dow Street, Dundee DD1 5EH, Scotland, U.K.
Organizational Affiliation: