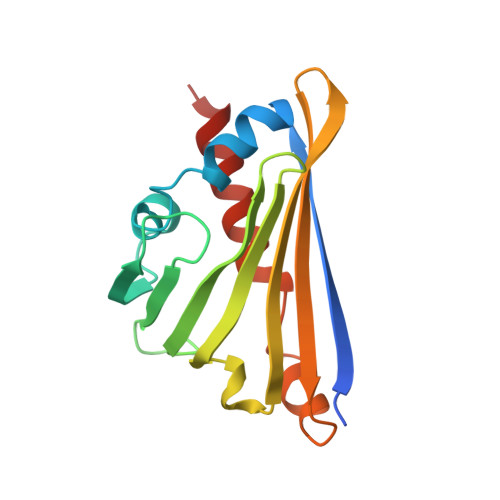
Structural Evidence for the Dopamine-First Mechanism of Norcoclaurine Synthase.
Lichman, B.R., Sula, A., Pesnot, T., Hailes, H.C., Ward, J.M., Keep, N.H.(2017) Biochemistry 56: 5274-5277
- PubMed: 28915025 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.7b00769
- Primary Citation Related Structures:
5N8Q, 5NON - PubMed Abstract:
Norcoclaurine synthase (NCS) is a Pictet-Spenglerase that catalyzes the first key step in plant benzylisoquinoline alkaloid metabolism, a compound family that includes bioactive natural products such as morphine. The enzyme has also shown great potential as a biocatalyst for the formation of chiral isoquinolines. Here we present new high-resolution X-ray crystallography data describing Thalictrum flavum NCS bound to a mechanism-inspired ligand. The structure supports two key features of the NCS "dopamine-first" mechanism: the binding of dopamine catechol to Lys-122 and the position of the carbonyl substrate binding site at the active site entrance. The catalytically vital residue Glu-110 occupies a previously unobserved ligand-bound conformation that may be catalytically significant. The potential roles of inhibitory binding and alternative amino acid conformations in the mechanism have also been revealed. This work significantly advances our understanding of the NCS mechanism and will aid future efforts to engineer the substrate scope and catalytic properties of this useful biocatalyst.
- Department of Biochemical Engineering, University College London , Gower Street, London WC1E 6BT, U.K.
Organizational Affiliation: