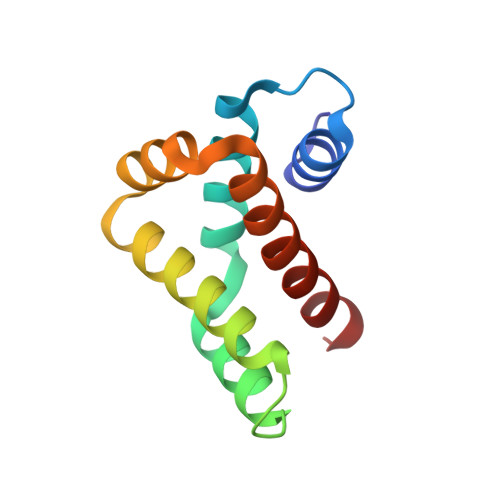
Structure and stability of the Human respiratory syncytial virus M2-1RNA-binding core domain reveals a compact and cooperative folding unit.
Molina, I.G., Josts, I., Almeida Hernandez, Y., Esperante, S., Salgueiro, M., Garcia Alai, M.M., de Prat-Gay, G., Tidow, H.(2018) Acta Crystallogr F Struct Biol Commun 74: 23-30
- PubMed: 29372904 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17017381
- Primary Citation Related Structures:
5NKX, 5NOH - PubMed Abstract:
Human syncytial respiratory virus is a nonsegmented negative-strand RNA virus with serious implications for respiratory disease in infants, and has recently been reclassified into a new family, Pneumoviridae. One of the main reasons for this classification is the unique presence of a transcriptional antiterminator, called M 2-1 . The puzzling mechanism of action of M 2-1 , which is a rarity among antiterminators in viruses and is part of the RNA polymerase complex, relies on dissecting the structure and function of this multidomain tetramer. The RNA-binding activity is located in a monomeric globular `core' domain, a high-resolution crystal structure of which is now presented. The structure reveals a compact domain which is superimposable on the full-length M 2-1 tetramer, with additional electron density for the C-terminal tail that was not observed in the previous models. Moreover, its folding stability was determined through chemical denaturation, which shows that the secondary and tertiary structure unfold concomitantly, which is indicative of a two-state equilibrium. These results constitute a further step in the understanding of this unique RNA-binding domain, for which there is no sequence or structural counterpart outside this virus family, in addition to its implications in transcription regulation and its likeliness as an antiviral target.
- Protein Structure-Function and Engineering Laboratory, Fundación Instituto Leloir and IIBBA-CONICET, Avenida Patricias Argentinas 435, C1405BWE Buenos Aires, Argentina.
Organizational Affiliation: