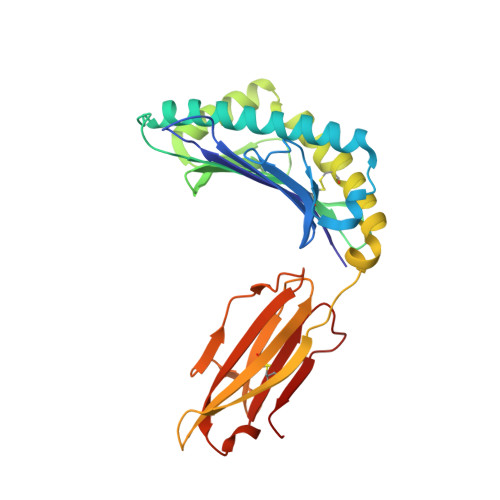
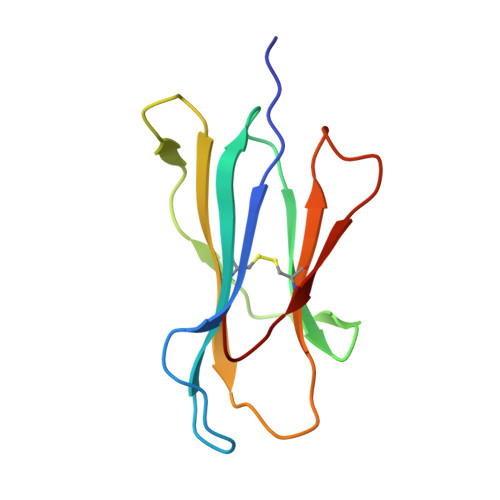
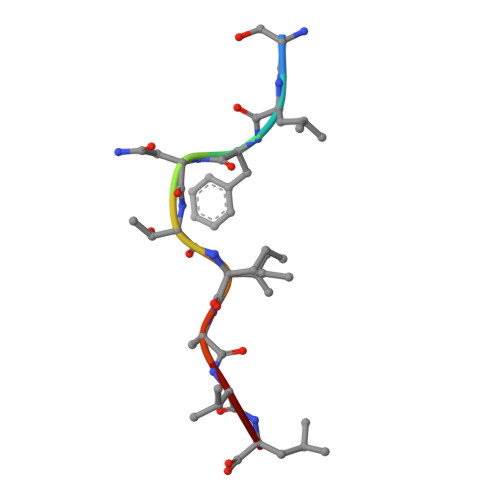
Dual Molecular Mechanisms Govern Escape at Immunodominant HLA A2-Restricted HIV Epitope.
Cole, D.K., Fuller, A., Dolton, G., Zervoudi, E., Legut, M., Miles, K., Blanchfield, L., Madura, F., Holland, C.J., Bulek, A.M., Bridgeman, J.S., Miles, J.J., Schauenburg, A.J.A., Beck, K., Evavold, B.D., Rizkallah, P.J., Sewell, A.K.(2017) Front Immunol 8: 1503-1503
- PubMed: 29209312
- DOI: https://doi.org/10.3389/fimmu.2017.01503
- Primary Citation Related Structures:
5NMD, 5NME, 5NMF, 5NMG, 5NMH, 5NMK - PubMed Abstract:
Serial accumulation of mutations to fixation in the SLYNTVATL (SL9) immunodominant, HIV p17 Gag-derived, HLA A2-restricted cytotoxic T lymphocyte epitope produce the SLFNTIAVL triple mutant "ultimate" escape variant. These mutations in solvent-exposed residues are believed to interfere with TCR recognition, although confirmation has awaited structural verification. Here, we solved a TCR co-complex structure with SL9 and the triple escape mutant to determine the mechanism of immune escape in this eminent system. We show that, in contrast to prevailing hypotheses, the main TCR contact residue is 4N and the dominant mechanism of escape is not via lack of TCR engagement. Instead, mutation of solvent-exposed residues in the peptide destabilise the peptide-HLA and reduce peptide density at the cell surface. These results highlight the extraordinary lengths that HIV employs to evade detection by high-affinity TCRs with a broad peptide-binding footprint and necessitate re-evaluation of this exemplar model of HIV TCR escape.
- Cardiff University School of Medicine, University Hospital, Heath Park, Cardiff, United Kingdom.
Organizational Affiliation: