New paradigms for understanding and step changes in treating active and chronic, persistent apicomplexan infections.
McPhillie, M., Zhou, Y., El Bissati, K., Dubey, J., Lorenzi, H., Capper, M., Lukens, A.K., Hickman, M., Muench, S., Verma, S.K., Weber, C.R., Wheeler, K., Gordon, J., Sanders, J., Moulton, H., Wang, K., Kim, T.K., He, Y., Santos, T., Woods, S., Lee, P., Donkin, D., Kim, E., Fraczek, L., Lykins, J., Esaa, F., Alibana-Clouser, F., Dovgin, S., Weiss, L., Brasseur, G., Wirth, D., Kent, M., Hood, L., Meunieur, B., Roberts, C.W., Hasnain, S.S., Antonyuk, S.V., Fishwick, C., McLeod, R.(2016) Sci Rep 6: 29179
- PubMed: 27412848 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep29179
- Primary Citation Related Structures:
5NMI - PubMed Abstract:
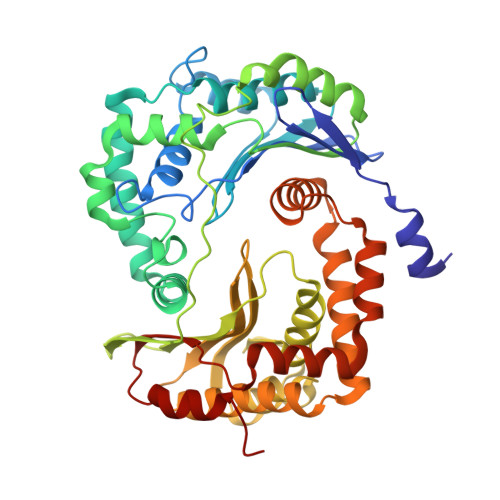
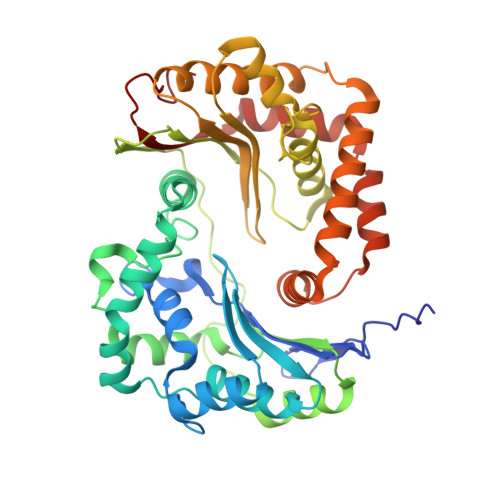
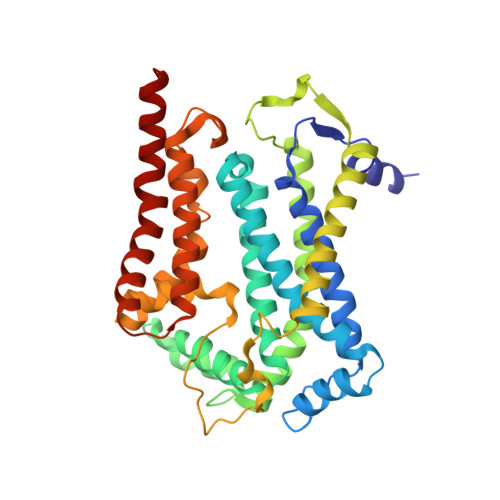
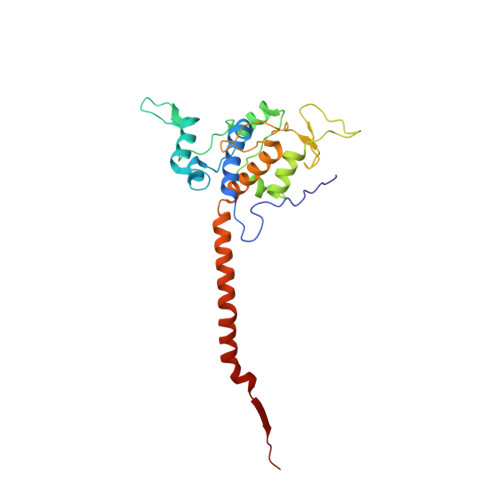
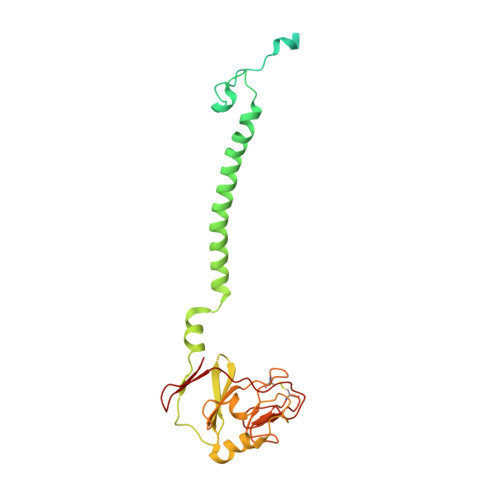
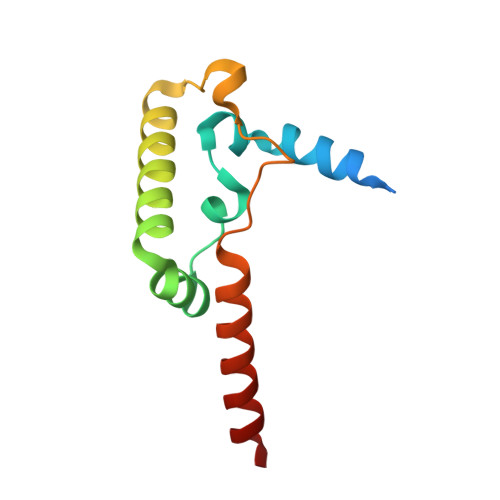
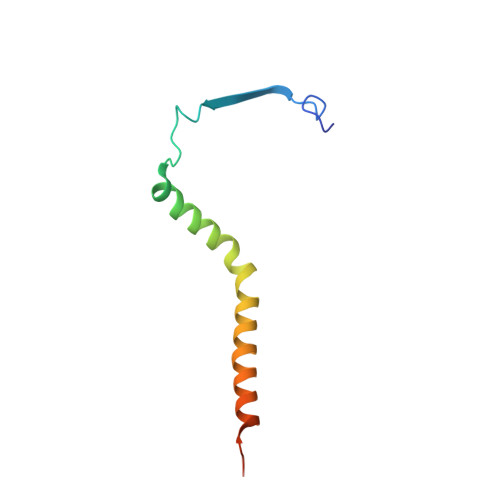
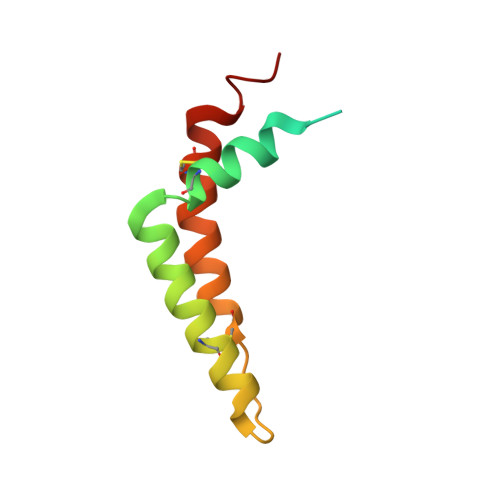
Toxoplasma gondii, the most common parasitic infection of human brain and eye, persists across lifetimes, can progressively damage sight, and is currently incurable. New, curative medicines are needed urgently. Herein, we develop novel models to facilitate drug development: EGS strain T. gondii forms cysts in vitro that induce oocysts in cats, the gold standard criterion for cysts. These cysts highly express cytochrome b. Using these models, we envisioned, and then created, novel 4-(1H)-quinolone scaffolds that target the cytochrome bc1 complex Qi site, of which, a substituted 5,6,7,8-tetrahydroquinolin-4-one inhibits active infection (IC50, 30 nM) and cysts (IC50, 4 μM) in vitro, and in vivo (25 mg/kg), and drug resistant Plasmodium falciparum (IC50, <30 nM), with clinically relevant synergy. Mutant yeast and co-crystallographic studies demonstrate binding to the bc1 complex Qi site. Our results have direct impact on improving outcomes for those with toxoplasmosis, malaria, and ~2 billion persons chronically infected with encysted bradyzoites.
- University of Leeds, Leeds, UK.
Organizational Affiliation: