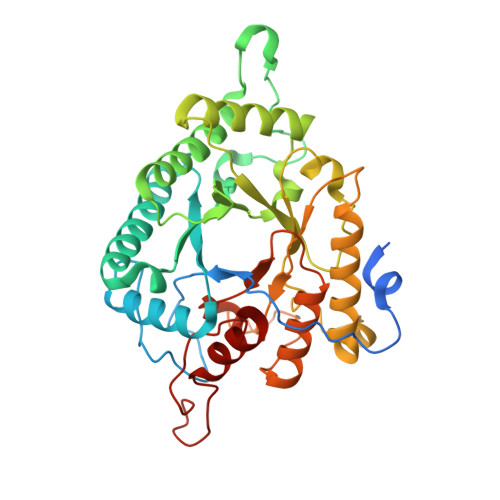
Crystal structure of a marine glycoside hydrolase family 99-related protein lacking catalytic machinery.
Robb, C.S., Mystkowska, A.A., Hehemann, J.H.(2017) Protein Sci 26: 2445-2450
- PubMed: 28884852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3291
- Primary Citation Related Structures:
5NGW - PubMed Abstract:
Algal polysaccharides of diverse structures are one of the most abundant carbon resources for heterotrophic, marine bacteria with coevolved digestive enzymes. A putative sulfo-mannan polysaccharide utilization locus, which is conserved in marine flavobacteria, contains an unusual GH99-like protein that lacks the conserved catalytic residues of glycoside hydrolase family 99. Using X-ray crystallography, we structurally characterized this protein from the marine flavobacterium Ochrovirga pacifica to help elucidate its molecular function. The structure reveals the absence of potential catalytic residues for polysaccharide hydrolysis, which-together with additional structural features-suggests this protein may be noncatalytic and involved in carbohydrate binding.
- Center for Marine Environmental Sciences, University of Bremen (MARUM), Bremen, 28359, Germany.
Organizational Affiliation: