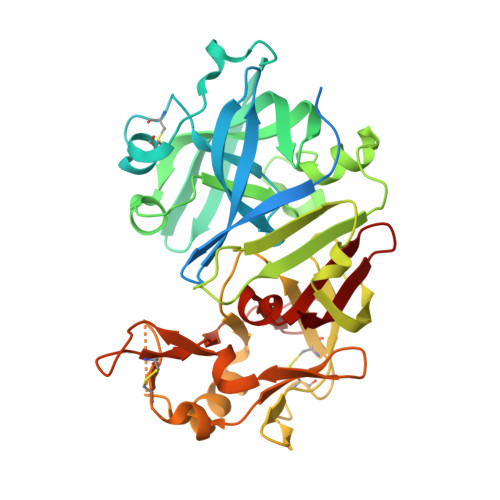
Functional and structural characterization of synthetic cardosin B-derived rennet.
Almeida, C.M., Manso, J.A., Figueiredo, A.C., Antunes, L., Cruz, R., Manadas, B., Bur, D., Pereira, P.J.B., Faro, C., Simoes, I.(2017) Appl Microbiol Biotechnol 101: 6951-6968
- PubMed: 28770303 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-017-8445-8
- Primary Citation Related Structures:
5NFG - PubMed Abstract:
The potential of using a synthetic cardosin-based rennet in cheese manufacturing was recently demonstrated with the development and optimization of production of a recombinant form of cardosin B in Kluyveromyces lactis. With the goal of providing a more detailed characterization of this rennet, we herein evaluate the impact of the plant-specific insert (PSI) on cardosin B secretion in this yeast, and provide a thorough analysis of the specificity requirements as well as the biochemical and structural properties of the isolated recombinant protease. We demonstrate that the PSI domain can be substituted by different linker sequences without substantially affecting protein secretion and milk clotting activity. However, the presence of small portions of the PSI results in dramatic reductions of secretion yields in this heterologous system. Kinetic characterization and specificity profiling results clearly suggest that synthetic cardosin B displays lower catalytic efficiency and is more sequence selective than native cardosin B. Elucidation of the structure of synthetic cardosin B confirms the canonical fold of an aspartic protease with the presence of two high mannose-type, N-linked glycan structures; however, there are some differences in the conformation of the flap region when compared to cardosin A. These subtle variations in catalytic properties and the more stringent substrate specificity of synthetic cardosin B help to explain the observed suitability of this rennet for cheese production.
- Biocant, Biotechnology Innovation Center, Parque Tecnológico de Cantanhede, Núcleo 4 Lote 8, 3060-197, Cantanhede, Portugal.
Organizational Affiliation: