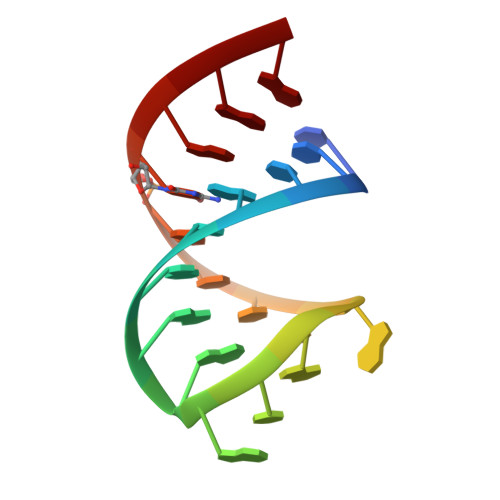
The Structure of the Guanidine-II Riboswitch.
Huang, L., Wang, J., Lilley, D.M.J.(2017) Cell Chem Biol 24: 695-702.e2
- PubMed: 28529131 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2017.05.014
- Primary Citation Related Structures:
5NDH, 5NDI, 5NEF, 5NEO, 5NEP, 5NEQ, 5NEX, 5NOM - PubMed Abstract:
The guanidine-II (mini-ykkC) riboswitch is the smallest of the guanidine-responsive riboswitches, comprising two stem loops of similar sequence. We have solved high-resolution crystal structures of both stem loops for the riboswitch from Gloeobacter violaceus. The stem loops have a strong propensity to dimerize by intimate loop-loop interaction. The dimerization creates specific binding pockets for two guanidine molecules, explaining their cooperative binding. Within the binding pockets the ligands are hydrogen bonded to a guanine at O6 and N7, and to successive backbone phosphates. Additionally they are each stacked upon a guanine nucleobase. One side of the pocket has an opening to the solvent, slightly lowering the specificity of ligand binding, and structures with bound methylguanidine, aminoguanidine, and agmatine show how this is possible.
- Cancer Research UK Nucleic Acid Structure Research Group, MSI/WTB Complex, The University of Dundee, Dow Street, Dundee DD1 5EH, UK.
Organizational Affiliation: