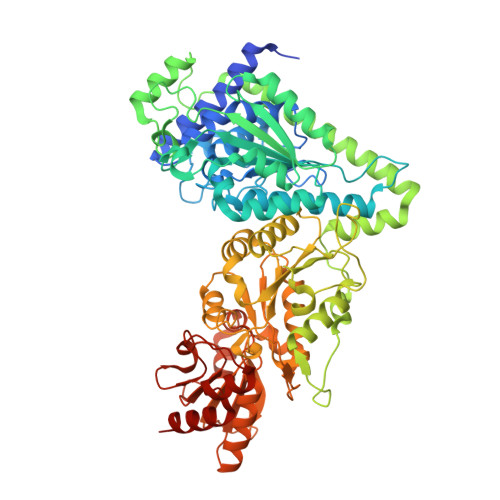
Structural basis for the magnesium-dependent activation of transketolase from Chlamydomonas reinhardtii.
Pasquini, M., Fermani, S., Tedesco, D., Sciabolini, C., Crozet, P., Naldi, M., Henri, J., Vothknecht, U., Bertucci, C., Lemaire, S.D., Zaffagnini, M., Francia, F.(2017) Biochim Biophys Acta 1861: 2132-2145
- PubMed: 28552632 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2017.05.021
- Primary Citation Related Structures:
5ND5, 5ND6 - PubMed Abstract:
In photosynthetic organisms, transketolase (TK) is involved in the Calvin-Benson cycle and participates to the regeneration of ribulose-5-phosphate. Previous studies demonstrated that TK catalysis is strictly dependent on thiamine pyrophosphate (TPP) and divalent ions such as Mg 2+ . TK from the unicellular green alga Chlamydomonas reinhardtii (CrTK) was recombinantly produced and purified to homogeneity. Biochemical properties of the CrTK enzyme were delineated by activity assays and its structural features determined by CD analysis and X-ray crystallography. CrTK is homodimeric and its catalysis depends on the reconstitution of the holo-enzyme in the presence of both TPP and Mg 2+ . Activity measurements and CD analysis revealed that the formation of fully active holo-CrTK is Mg 2+ -dependent and proceeds with a slow kinetics. The 3D-structure of CrTK without cofactors (CrTK apo ) shows that two portions of the active site are flexible and disordered while they adopt an ordered conformation in the holo-form. Oxidative treatments revealed that Mg 2+ participates in the redox control of CrTK by changing its propensity to be inactivated by oxidation. Indeed, the activity of holo-form is unaffected by oxidation whereas CrTK in the apo-form or reconstituted with the sole TPP show a strong sensitivity to oxidative inactivation. These evidences indicate that Mg 2+ is fundamental to allow gradual conformational arrangements suited for optimal catalysis. Moreover, Mg 2+ is involved in the control of redox sensitivity of CrTK. The importance of Mg 2+ in the functionality and redox sensitivity of CrTK is correlated to light-dependent fluctuations of Mg 2+ in chloroplasts.
- Department of Pharmacy and Biotechnology, University of Bologna, Bologna, Italy.
Organizational Affiliation: