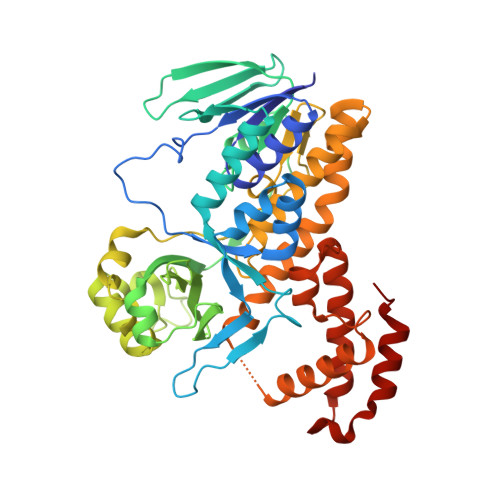
Structural and mechanistic basis of differentiated inhibitors of the acute pancreatitis target kynurenine-3-monooxygenase.
Hutchinson, J.P., Rowland, P., Taylor, M.R.D., Christodoulou, E.M., Haslam, C., Hobbs, C.I., Holmes, D.S., Homes, P., Liddle, J., Mole, D.J., Uings, I., Walker, A.L., Webster, S.P., Mowat, C.G., Chung, C.W.(2017) Nat Commun 8: 15827-15827
- PubMed: 28604669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15827
- Primary Citation Related Structures:
5NA5, 5NAB, 5NAE, 5NAG, 5NAH, 5NAK - PubMed Abstract:
Kynurenine-3-monooxygenase (KMO) is a key FAD-dependent enzyme of tryptophan metabolism. In animal models, KMO inhibition has shown benefit in neurodegenerative diseases such as Huntington's and Alzheimer's. Most recently it has been identified as a target for acute pancreatitis multiple organ dysfunction syndrome (AP-MODS); a devastating inflammatory condition with a mortality rate in excess of 20%. Here we report and dissect the molecular mechanism of action of three classes of KMO inhibitors with differentiated binding modes and kinetics. Two novel inhibitor classes trap the catalytic flavin in a previously unobserved tilting conformation. This correlates with picomolar affinities, increased residence times and an absence of the peroxide production seen with previous substrate site inhibitors. These structural and mechanistic insights culminated in GSK065(C1) and GSK366(C2), molecules suitable for preclinical evaluation. Moreover, revising the repertoire of flavin dynamics in this enzyme class offers exciting new opportunities for inhibitor design.
- Platform Technologies and Science, GlaxoSmithKline, Stevenage SG1 2NY, UK.
Organizational Affiliation: