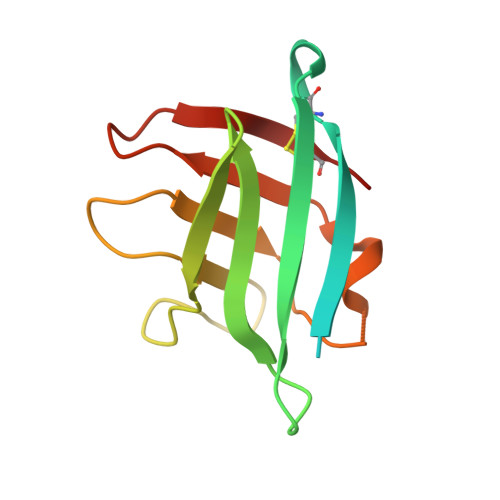
Structure of the Neisseria Adhesin Complex Protein (ACP) and its role as a novel lysozyme inhibitor.
Humbert, M.V., Awanye, A.M., Lian, L.Y., Derrick, J.P., Christodoulides, M.(2017) PLoS Pathog 13: e1006448-e1006448
- PubMed: 28662181 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1006448
- Primary Citation Related Structures:
5MY7 - PubMed Abstract:
Pathogenic and commensal Neisseria species produce an Adhesin Complex Protein, which was first characterised in Neisseria meningitidis (Nm) as a novel surface-exposed adhesin with vaccine potential. In the current study, the crystal structure of a recombinant (r)Nm-ACP Type I protein was determined to 1.4 Å resolution: the fold resembles an eight-stranded β-barrel, stabilized by a disulphide bond between the first (Cys38) and last (Cys121) β-strands. There are few main-chain hydrogen bonds linking β4-β5 and β8-β1, so the structure divides into two four-stranded anti-parallel β-sheets (β1-β4 and β5-β8). The computed surface electrostatic charge distribution showed that the β1-β4 sheet face is predominantly basic, whereas the β5-β8 sheet is apolar, apart from the loop between β4 and β5. Concentrations of rNm-ACP and rNeisseria gonorrhoeae-ACP proteins ≥0.25 μg/ml significantly inhibited by ~80-100% (P<0.05) the in vitro activity of human lysozyme (HL) over 24 h. Specificity was demonstrated by the ability of murine anti-Neisseria ACP sera to block ACP inhibition and restore HL activity. ACP expression conferred tolerance to HL activity, as demonstrated by significant 3-9 fold reductions (P<0.05) in the growth of meningococcal and gonococcal acp gene knock-out mutants in the presence of lysozyme. In addition, wild-type Neisseria lactamica treated with purified ACP-specific rabbit IgG antibodies showed similar fold reductions in bacterial growth, compared with untreated bacteria (P<0.05). Nm-ACPI is structurally similar to the MliC/PliC protein family of lysozyme inhibitors. However, Neisseria ACP proteins show <20% primary sequence similarity with these inhibitors and do not share any conserved MliC/PliC sequence motifs associated with lysozyme recognition. These observations suggest that Neisseria ACP adopts a different mode of lysozyme inhibition and that the ability of ACP to inhibit lysozyme activity could be important for host colonization by both pathogenic and commensal Neisseria organisms. Thus, ACP represents a dual target for developing Neisseria vaccines and drugs to inhibit host-pathogen interactions.
- Neisseria Research, Molecular Microbiology, Academic Unit of Clinical and Experimental Sciences, Sir Henry Wellcome Laboratories, University of Southampton Faculty of Medicine, Southampton, United Kingdom.
Organizational Affiliation: