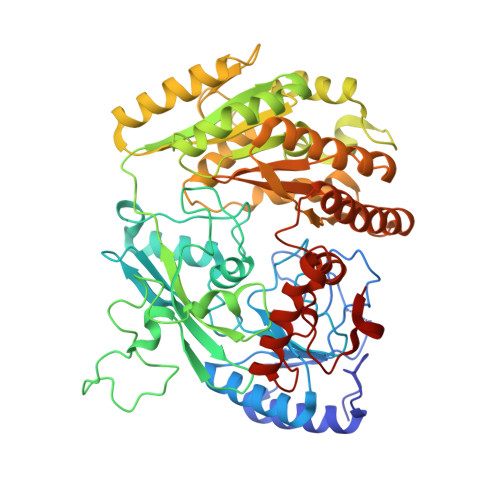
Two tyrosine residues, Tyr-108 and Tyr-503, are responsible for the deprotonation of phenolic substrates in vanillyl-alcohol oxidase.
Ewing, T.A., Nguyen, Q.T., Allan, R.C., Gygli, G., Romero, E., Binda, C., Fraaije, M.W., Mattevi, A., van Berkel, W.J.H.(2017) J Biological Chem 292: 14668-14679
- PubMed: 28717004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.778449
- Primary Citation Related Structures:
5MXJ, 5MXU - PubMed Abstract:
A number of oxidoreductases from the VAO/ para -cresol methylhydroxylase flavoprotein family catalyze the oxidation of para -substituted phenols. One of the best-studied is vanillyl-alcohol oxidase (VAO) from the fungus Penicillium simplicissimum For oxidation of phenols by VAO to occur, they must first be bound in the active site of the enzyme in their phenolate anion form. The crystal structure of VAO reveals that two tyrosine residues, Tyr-108 and Tyr-503, are positioned to facilitate this deprotonation. To investigate their role in catalysis, we created three VAO variants, Y108F, Y503F, and Y108F/Y503F, and studied their biochemical properties. Steady-state kinetics indicated that the presence of at least one of the tyrosine residues is essential for efficient catalysis by VAO. Stopped-flow kinetics revealed that the reduction of VAO by chavicol or vanillyl alcohol occurs at two different rates: k obs1 , which corresponds to its reaction with the deprotonated form of the substrate, and k obs2 , which corresponds to its reaction with the protonated form of the substrate. In Y108F, Y503F, and Y108F/Y503F, the relative contribution of k obs2 to the reduction is larger than in wild-type VAO, suggesting deprotonation is impaired in these variants. Binding studies disclosed that the competitive inhibitor isoeugenol is predominantly in its deprotonated form when bound to wild-type VAO, but predominantly in its protonated form when bound to the variants. These results indicate that Tyr-108 and Tyr-503 are responsible for the activation of substrates in VAO, providing new insights into the catalytic mechanism of VAO and related enzymes that oxidize para -substituted phenols.
- From the Laboratory of Biochemistry, Wageningen University & Research, Stippeneng 4, 6708 WE Wageningen, The Netherlands.
Organizational Affiliation: