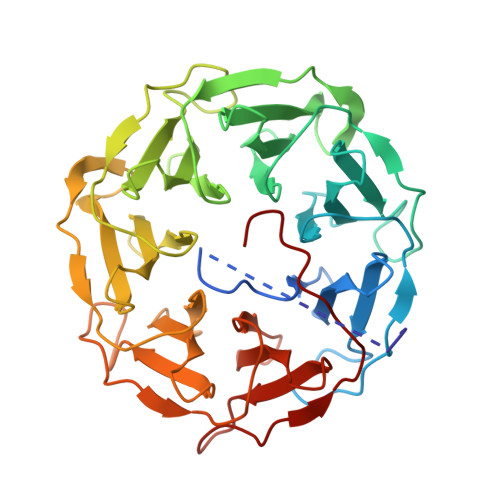
Characterization of novel bangle lectin from Photorhabdus asymbiotica with dual sugar-binding specificity and its effect on host immunity.
Jancarikova, G., Houser, J., Dobes, P., Demo, G., Hyrsl, P., Wimmerova, M.(2017) PLoS Pathog 13: e1006564-e1006564
- PubMed: 28806750 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1006564
- Primary Citation Related Structures:
5MXE, 5MXF, 5MXG, 5MXH - PubMed Abstract:
Photorhabdus asymbiotica is one of the three recognized species of the Photorhabdus genus, which consists of gram-negative bioluminescent bacteria belonging to the family Morganellaceae. These bacteria live in a symbiotic relationship with nematodes from the genus Heterorhabditis, together forming a complex that is highly pathogenic for insects. Unlike other Photorhabdus species, which are strictly entomopathogenic, P. asymbiotica is unique in its ability to act as an emerging human pathogen. Analysis of the P. asymbiotica genome identified a novel fucose-binding lectin designated PHL with a strong sequence similarity to the recently described P. luminescens lectin PLL. Recombinant PHL exhibited high affinity for fucosylated carbohydrates and the unusual disaccharide 3,6-O-Me2-Glcβ1-4(2,3-O-Me2)Rhaα-O-(p-C6H4)-OCH2CH2NH2 from Mycobacterium leprae. Based on its crystal structure, PHL forms a seven-bladed β-propeller assembling into a homo-dimer with an inter-subunit disulfide bridge. Investigating complexes with different ligands revealed the existence of two sets of binding sites per monomer-the first type prefers l-fucose and its derivatives, whereas the second type can bind d-galactose. Based on the sequence analysis, PHL could contain up to twelve binding sites per monomer. PHL was shown to interact with all types of red blood cells and insect haemocytes. Interestingly, PHL inhibited the production of reactive oxygen species induced by zymosan A in human blood and antimicrobial activity both in human blood, serum and insect haemolymph. Concurrently, PHL increased the constitutive level of oxidants in the blood and induced melanisation in haemolymph. Our results suggest that PHL might play a crucial role in the interaction of P. asymbiotica with both human and insect hosts.
- Central European Institute of Technology (CEITEC), Masaryk University, Brno, Czech Republic.
Organizational Affiliation: