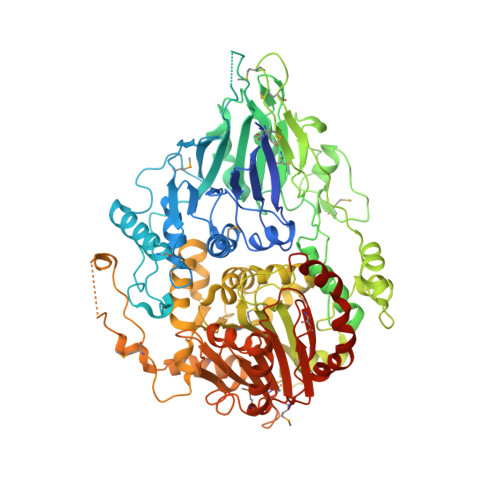
The crystal structure of Trz1, the long form RNase Z from yeast.
Ma, M., Li de la Sierra-Gallay, I., Lazar, N., Pellegrini, O., Durand, D., Marchfelder, A., Condon, C., van Tilbeurgh, H.(2017) Nucleic Acids Res 45: 6209-6216
- PubMed: 28379452 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx216
- Primary Citation Related Structures:
5MTZ - PubMed Abstract:
tRNAs are synthesized as precursor RNAs that have to undergo processing steps to become functional. Yeast Trz1 is a key endoribonuclease involved in the 3΄ maturation of tRNAs in all domains of life. It is a member of the β-lactamase family of RNases, characterized by an HxHxDH sequence motif involved in coordination of catalytic Zn-ions. The RNase Z family consists of two subfamilies: the short (250-400 residues) and the long forms (about double in size). Short form RNase Z enzymes act as homodimers: one subunit embraces tRNA with a protruding arm, while the other provides the catalytic site. The long form is thought to contain two fused β-lactamase domains within a single polypeptide. Only structures of short form RNase Z enzymes are known. Here we present the 3.1 Å crystal structure of the long-form Trz1 from Saccharomyces cerevisiae. Trz1 is organized into two β-lactamase domains connected by a long linker. The N-terminal domain has lost its catalytic residues, but retains the long flexible arm that is important for tRNA binding, while it is the other way around in the C-terminal domain. Trz1 likely evolved from a duplication and fusion of the gene encoding the monomeric short form RNase Z.
- Institute for Integrative Biology of the Cell (I2BC), CEA, CNRS UMR 9198, University of Paris-Sud, Université Paris-Saclay, 91198 Gif sur Yvette Cedex, France.
Organizational Affiliation: