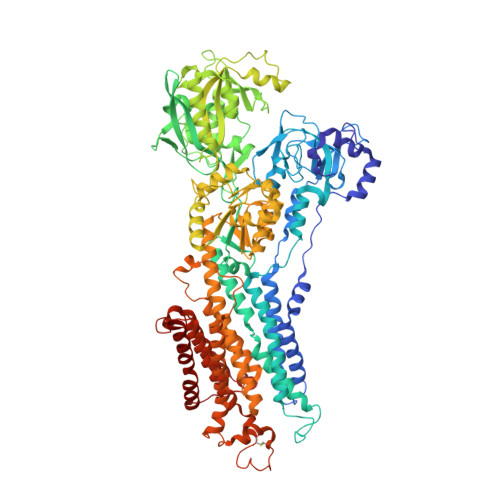
Structures of the heart specific SERCA2a Ca2+-ATPase.
Sitsel, A., De Raeymaecker, J., Drachmann, N.D., Derua, R., Smaardijk, S., Andersen, J.L., Vandecaetsbeek, I., Chen, J., De Maeyer, M., Waelkens, E., Olesen, C., Vangheluwe, P., Nissen, P.(2019) EMBO J 38
- PubMed: 30777856 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2018100020
- Primary Citation Related Structures:
5MPM, 6HXB - PubMed Abstract:
The sarcoplasmic/endoplasmic reticulum Ca 2+ -ATPase 2a (SERCA2a) performs active reuptake of cytoplasmic Ca 2+ and is a major regulator of cardiac muscle contractility. Dysfunction or dysregulation of SERCA2a is associated with heart failure, while restoring its function is considered as a therapeutic strategy to restore cardiac performance. However, its structure has not yet been determined. Based on native, active protein purified from pig ventricular muscle, we present the first crystal structures of SERCA2a, determined in the CPA-stabilized E2-AlF4- form (3.3 Å) and the Ca 2+ -occluded [Ca 2 ]E1-AMPPCP form (4.0 Å). The structures are similar to the skeletal muscle isoform SERCA1a pointing to a conserved mechanism. We seek to explain the kinetic differences between SERCA1a and SERCA2a. We find that several isoform-specific residues are acceptor sites for post-translational modifications. In addition, molecular dynamics simulations predict that isoform-specific residues support distinct intramolecular interactions in SERCA2a and SERCA1a. Our experimental observations further indicate that isoform-specific intramolecular interactions are functionally relevant, and may explain the kinetic differences between SERCA2a and SERCA1a.
- Department of Molecular Biology and Genetics, Aarhus University, Aarhus, Denmark.
Organizational Affiliation: