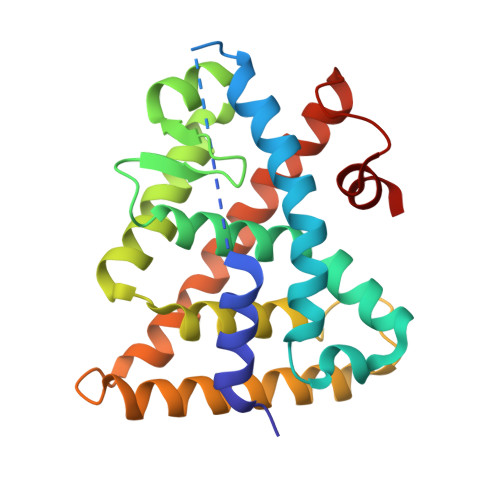
Ligand Dependent Switch from RXR Homo- to RXR-NURR1 Heterodimerization.
Scheepstra, M., Andrei, S.A., de Vries, R.M.J.M., Meijer, F.A., Ma, J.N., Burstein, E.S., Olsson, R., Ottmann, C., Milroy, L.G., Brunsveld, L.(2017) ACS Chem Neurosci 8: 2065-2077
- PubMed: 28691794 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschemneuro.7b00216
- Primary Citation Related Structures:
5MJ5, 5MK4, 5MKJ, 5MKU, 5MMW - PubMed Abstract:
Retinoid X receptors (RXRs) play key roles in many physiological processes in both the periphery and central nervous system. In addition, RXRs form heterodimers with other nuclear receptors to exert their physiological effects. The nuclear receptor related 1 protein (NURR1) is particularly interesting because of its role in promoting differentiation and survival of dopamine neurons. However, only a small number of RXR-heterodimer selective modulators are available, with limited chemical diversity. This work describes the synthesis, biochemical evaluation, and structural elucidation of a novel series of RXR ligands with strongly biased interactions with RXRα-NURR1 heterodimers. Targeted modifications to the small molecule biaryl scaffold caused local RXRα side-chain disturbances and displacement of secondary structural elements upon ligand binding. This resulted in the repositioning of protein helices in the heterodimer interface of RXRα, alterations in homo- versus heterodimer formation, and modulation of activation function 2 (AF2). The data provide a rationale for the design of RXR ligands consisting of a highly conserved hydrophilic region, strongly contributing to the ligand affinity, and a variable hydrophobic region, which efficiently probes the effects of structural changes at the level of the ligand on co-regulator recruitment or the RXRα-NURR1 dimerization interface.
- Department of Biomedical Engineering and Institute for Complex Molecular Systems, Laboratory of Chemical Biology, Technische Universiteit Eindhoven , Den Dolech 2, 5612 AZ Eindhoven, The Netherlands.
Organizational Affiliation: