Mapping the affinity landscape of Thrombin-binding aptamers on 2 F-ANA/DNA chimeric G-Quadruplex microarrays.
Lietard, J., Abou Assi, H., Gomez-Pinto, I., Gonzalez, C., Somoza, M.M., Damha, M.J.(2017) Nucleic Acids Res 45: 1619-1632
- PubMed: 28100695 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkw1357
- Primary Citation Related Structures:
5MJX - PubMed Abstract:
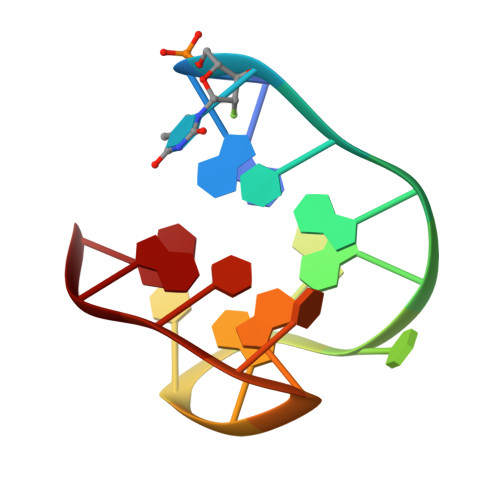
In situ fabricated nucleic acids microarrays are versatile and very high-throughput platforms for aptamer optimization and discovery, but the chemical space that can be probed against a given target has largely been confined to DNA, while RNA and non-natural nucleic acid microarrays are still an essentially uncharted territory. 2΄-Fluoroarabinonucleic acid (2΄F-ANA) is a prime candidate for such use in microarrays. Indeed, 2΄F-ANA chemistry is readily amenable to photolithographic microarray synthesis and its potential in high affinity aptamers has been recently discovered. We thus synthesized the first microarrays containing 2΄F-ANA and 2΄F-ANA/DNA chimeric sequences to fully map the binding affinity landscape of the TBA1 thrombin-binding G-quadruplex aptamer containing all 32 768 possible DNA-to-2΄F-ANA mutations. The resulting microarray was screened against thrombin to identify a series of promising 2΄F-ANA-modified aptamer candidates with Kds significantly lower than that of the unmodified control and which were found to adopt highly stable, antiparallel-folded G-quadruplex structures. The solution structure of the TBA1 aptamer modified with 2΄F-ANA at position T3 shows that fluorine substitution preorganizes the dinucleotide loop into the proper conformation for interaction with thrombin. Overall, our work strengthens the potential of 2΄F-ANA in aptamer research and further expands non-genomic applications of nucleic acids microarrays.
- Institute of Inorganic Chemistry, Faculty of Chemistry, University of Vienna, Althanstraße 14 (UZA II), 1090 Vienna, Austria.
Organizational Affiliation: