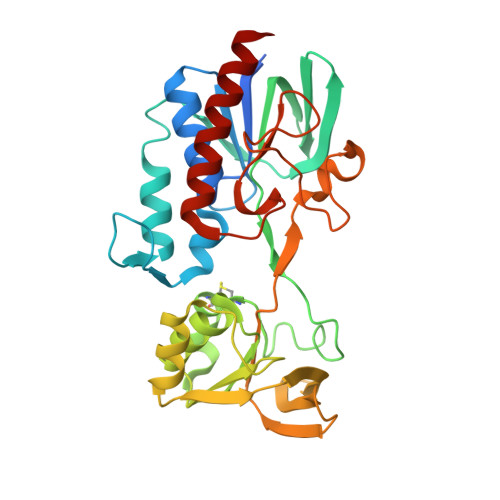
The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Skjoldager, N., Blanner Bang, M., Rykr, M., Bjornberg, O., Davies, M.J., Svensson, B., Harris, P., Hagglund, P.(2017) Sci Rep 7: 46282-46282
- PubMed: 28397795
- DOI: https://doi.org/10.1038/srep46282
- Primary Citation Related Structures:
5MH4, 5MIP, 5MIQ, 5MIR, 5MIS, 5MIT, 5MJK - PubMed Abstract:
The NADPH-dependent homodimeric flavoenzyme thioredoxin reductase (TrxR) provides reducing equivalents to thioredoxin, a key regulator of various cellular redox processes. Crystal structures of photo-inactivated thioredoxin reductase (TrxR) from the Gram-positive bacterium Lactococcus lactis have been determined. These structures reveal novel molecular features that provide further insight into the mechanisms behind the sensitivity of this enzyme toward visible light. We propose that a pocket on the si-face of the isoalloxazine ring accommodates oxygen that reacts with photo-excited FAD generating superoxide and a flavin radical that oxidize the isoalloxazine ring C7α methyl group and a nearby tyrosine residue. This tyrosine and key residues surrounding the oxygen pocket are conserved in enzymes from related bacteria, including pathogens such as Staphylococcus aureus. Photo-sensitivity may thus be a widespread feature among bacterial TrxR with the described characteristics, which affords applications in clinical photo-therapy of drug-resistant bacteria.
- Department of Biotechnology and Biomedicine, Technical University of Denmark, DK-2800 Kgs. Lyngby, Denmark.
Organizational Affiliation: