Structure and Catalytic Mechanism of a Bacterial Friedel-Crafts Acylase.
Pavkov-Keller, T., Schmidt, N.G., Zadlo-Dobrowolska, A., Kroutil, W., Gruber, K.(2019) Chembiochem 20: 88-95
- PubMed: 30318713 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cbic.201800462
- Primary Citation Related Structures:
5M3K, 5MG5 - PubMed Abstract:
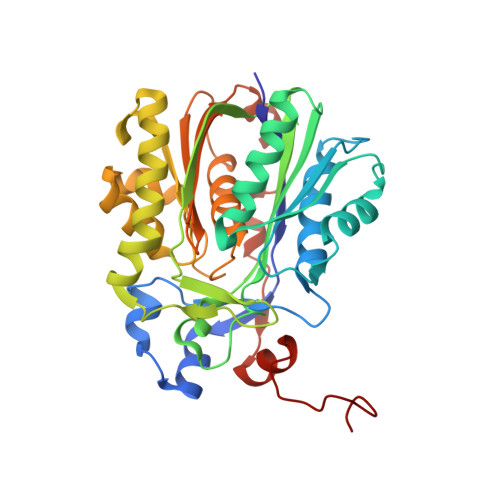
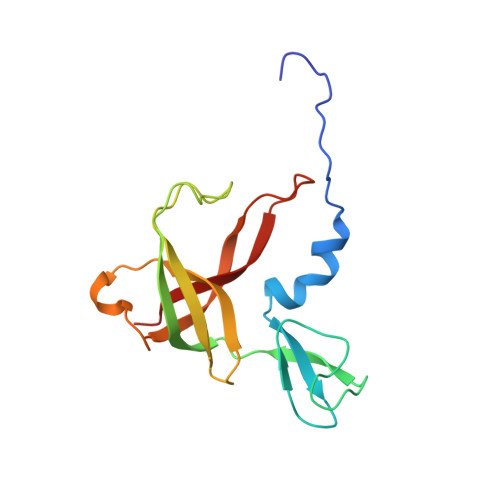
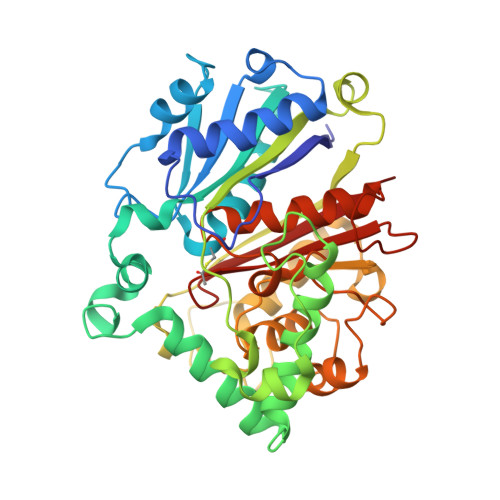
C-C bond-forming reactions are key transformations for setting up the carbon frameworks of organic compounds. In this context, Friedel-Crafts acylation is commonly used for the synthesis of aryl ketones, which are common motifs in many fine chemicals and natural products. A bacterial multicomponent acyltransferase from Pseudomonas protegens (PpATase) catalyzes such Friedel-Crafts C-acylation of phenolic substrates in aqueous solution, reaching up to >99 % conversion without the need for CoA-activated reagents. We determined X-ray crystal structures of the native and ligand-bound complexes. This multimeric enzyme consists of three subunits: PhlA, PhlB, and PhlC, arranged in a Phl(A 2 C 2 ) 2 B 4 composition. The structure of a reaction intermediate obtained from crystals soaked with the natural substrate 1-(2,4,6-trihydroxyphenyl)ethanone together with site-directed mutagenesis studies revealed that only residues from the PhlC subunits are involved in the acyl transfer reaction, with Cys88 very likely playing a significant role during catalysis. These structural and mechanistic insights form the basis of further enzyme engineering efforts directed towards enhancing the substrate scope of this enzyme.
- Austrian Centre of Industrial Biotechnology (ACIB), Petersgasse 14, 8010, Graz, Austria.
Organizational Affiliation: