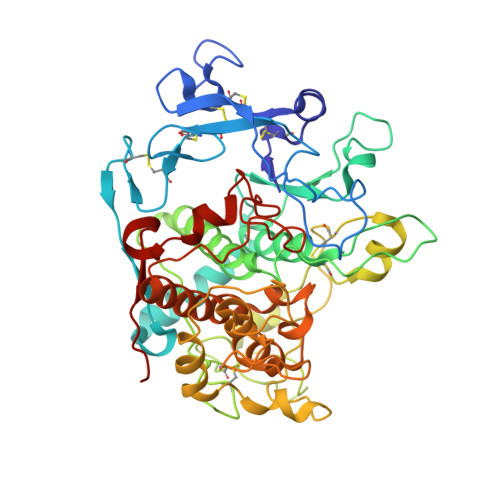
Structure of Human Tyrosinase Related Protein 1 Reveals a Binuclear Zinc Active Site Important for Melanogenesis.
Lai, X., Wichers, H.J., Soler-Lopez, M., Dijkstra, B.W.(2017) Angew Chem Int Ed Engl 56: 9812-9815
- PubMed: 28661582
- DOI: https://doi.org/10.1002/anie.201704616
- Primary Citation of Related Structures:
5M8L, 5M8M, 5M8N, 5M8O, 5M8P, 5M8Q, 5M8R, 5M8T - PubMed Abstract:
Tyrosinase-related protein 1 (TYRP1) is one of three tyrosinase-like glycoenzymes in human melanocytes that are key to the production of melanin, the compound responsible for the pigmentation of skin, eye, and hair. Difficulties with producing these enzymes in pure form have hampered the understanding of their activity and the effect of mutations that cause albinism and pigmentation disorders. Herein we show that the typical tyrosinase-like subdomain of TYRP1 contains two zinc ions in the active site instead of copper ions as found in tyrosinases, which explains why TYRP1 does not exhibit tyrosinase redox activity. In addition, the structures reveal for the first time that the Cys-rich subdomain, which is unique to vertebrate melanogenic proteins, has an epidermal growth factor-like fold and is tightly associated with the tyrosinase subdomain. Our structures suggest that most albinism-related mutations of TYRP1 affect its stability or activity.
- Laboratory of Biophysical Chemistry, University of Groningen, Nijenborgh 7, 9747 AG, Groningen, The Netherlands.
Organizational Affiliation: