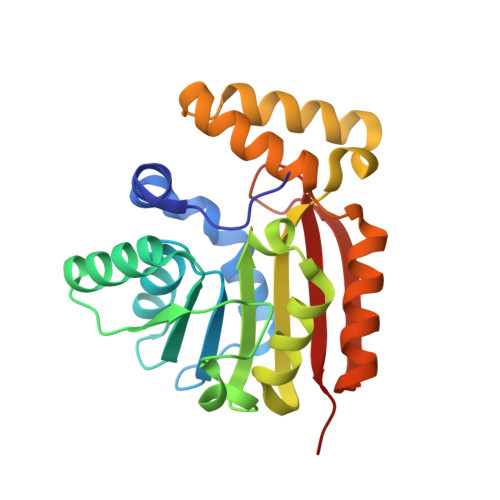
Crystal Structure and Catalytic Mechanism of CouO, a Versatile C-Methyltransferase from Streptomyces rishiriensis.
Pavkov-Keller, T., Steiner, K., Faber, M., Tengg, M., Schwab, H., Gruber-Khadjawi, M., Gruber, K.(2017) PLoS One 12: e0171056-e0171056
- PubMed: 28152088 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0171056
- Primary Citation Related Structures:
5M58 - PubMed Abstract:
Friedel-Crafts alkylation of aromatic systems is a classic reaction in organic chemistry, for which regiospecific mono-alkylation, however, is generally difficult to achieve. In nature, methyltransferases catalyze the addition of methyl groups to a wide range of biomolecules thereby modulating the physico-chemical properties of these compounds. Specifically, S-adenosyl-L-methionine dependent C-methyltransferases possess a high potential to serve as biocatalysts in environmentally benign organic syntheses. Here, we report on the high resolution crystal structure of CouO, a C-methyltransferase from Streptomyces rishiriensis involved in the biosynthesis of the antibiotic coumermycin A1. Through molecular docking calculations, site-directed mutagenesis and the comparison with homologous enzymes we identified His120 and Arg121 as key functional residues for the enzymatic activity of this group of C-methyltransferases. The elucidation of the atomic structure and the insight into the catalytic mechanism provide the basis for the (semi)-rational engineering of the enzyme in order to increase the substrate scope as well as to facilitate the acceptance of SAM-analogues as alternative cofactors.
- Institute of Molecular Biosciences, University of Graz, Graz, Austria.
Organizational Affiliation: