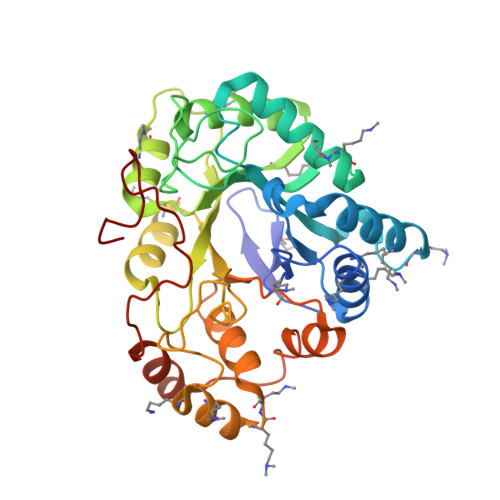
Structural basis for the inhibition of AKR1B10 by the C3 brominated TTNPB derivative UVI2008.
Ruiz, F.X., Crespo, I., Alvarez, S., Porte, S., Gimenez-Dejoz, J., Cousido-Siah, A., Mitschler, A., de Lera, A.R., Pares, X., Podjarny, A., Farres, J.(2017) Chem Biol Interact 276: 174-181
- PubMed: 28161411 Search on PubMed
- DOI: https://doi.org/10.1016/j.cbi.2017.01.026
- Primary Citation Related Structures:
5M2F - PubMed Abstract:
UVI2008, a retinoic acid receptor (RAR) β/γ agonist originated from C3 bromine addition to the parent RAR pan-agonist 4-[(E)-2-(5,6,7,8-tetrahydro-5,5,8,8-tetramethyl-2-naphthalenyl)-1-propenyl]benzoic acid (TTNPB), is also a selective inhibitor of aldo-keto reductase family member 1B10 (AKR1B10). Thus, it might become a lead drug for the design of compounds targeting both activities, as an AKR1B10 inhibitor and RAR agonist, which could constitute a novel therapeutic approach against cancer and skin-related diseases. Herein, the X-ray structure of the methylated Lys125Arg/Val301Leu AKR1B10 (i.e. AKME2MU) holoenzyme in complex with UVI2008 was determined at 1.5 Å resolution, providing an explanation for UVI2008 selectivity against AKR1B10 (IC 50 = 6.1 μM) over the closely related aldose reductase (AR, IC 50 = 70 μM). The carboxylic acid group of UVI2008 is located in the anion-binding pocket, at hydrogen-bond distance of catalytically important residues Tyr49 and His111. The inhibitor bromine atom can only fit in the wider active site of AKR1B10, mainly because of the native Trp112 side-chain orientation, not possible in AR. In AKR1B10, Trp112 native conformation, and thus UVI2008 binding, is facilitated through interaction with Gln114. IC 50 analysis of the corresponding Thr113Gln mutant in AR confirmed this hypothesis. The elucidation of the binding mode of UVI2008 to AKR1B10, along with the previous studies on the retinoid specificity of AKR1B10 and on the stilbene retinoid scaffold conforming UVI2008, could indeed be used to foster the drug design of bifunctional antiproliferative compounds.
- Institut de Génétique et de Biologie Moléculaire et Cellulaire, 1 rue Laurent Fries, 67404 Illkirch Cedex, France. Electronic address: fxavier.ruiz@gmail.com.
Organizational Affiliation: