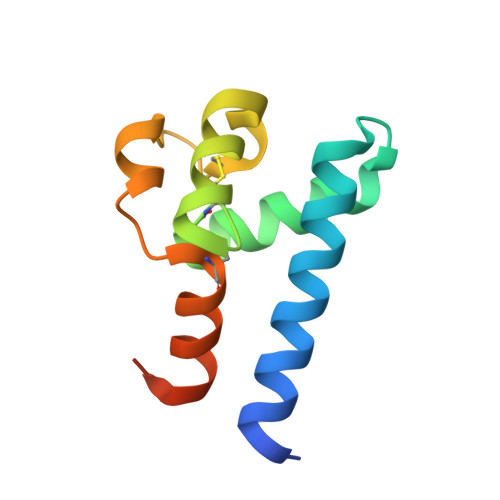
Mechanism of Structural Tuning of the Hepatitis C Virus Human Cellular Receptor CD81 Large Extracellular Loop.
Cunha, E.S., Sfriso, P., Rojas, A.L., Roversi, P., Hospital, A., Orozco, M., Abrescia, N.G.(2017) Structure 25: 53-65
- PubMed: 27916518 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2016.11.003
- Primary Citation Related Structures:
5M2C, 5M33, 5M3D, 5M3T, 5M4R - PubMed Abstract:
Hepatitis C virus (HCV) enters into human hepatocytes via tetraspanin hCD81. HCV glycoprotein E2 recognizes the "head" subdomain of the large extracellular loop (LEL) of CD81 (hCD81 LEL ), but the precise mechanism of virus cell attachment and entry remains elusive. Here, by combining the structural analysis of a conspicuous number of crystallized CD81 LEL molecules with molecular dynamics simulations, we show that the conformational plasticity of the hCD81 LEL head subdomain is a molecular property of the receptor. The observed closed, intermediate, and open conformations of the head subdomain provide distinct binding platforms. Simulations at pH 7.4 and 4.0 indicate that this dynamism is pH modulated. The crystallized double conformation of the disulfide bridge C157-C175 at the base of the head subdomain identifies this bond as the molecular zipper of the plasticity of hCD81 LEL . We propose that this conformational dependence of hCD81 LEL , which is finely tuned by pH and redox conditions, enables the virus-receptor interactions to diversely re-engage at endosomal conditions.
- Structural Biology Unit, CIC bioGUNE, CIBERehd, Derio 48160, Spain.
Organizational Affiliation: