Tools for the rational design of bivalent microtubule-targeting drugs.
Marangon, J., Christodoulou, M.S., Casagrande, F.V., Tiana, G., Dalla Via, L., Aliverti, A., Passarella, D., Cappelletti, G., Ricagno, S.(2016) Biochem Biophys Res Commun 479: 48-53
- PubMed: 27613098 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.09.022
- Primary Citation Related Structures:
5LP6 - PubMed Abstract:
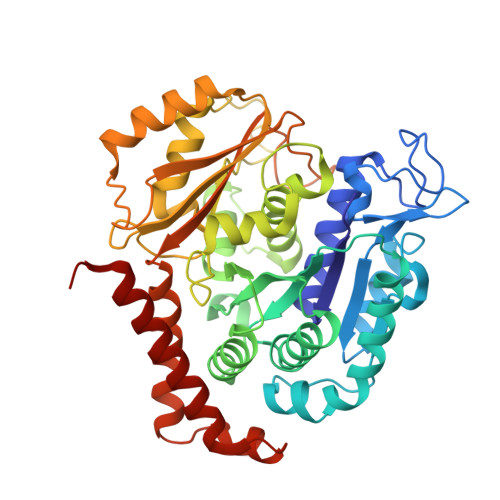
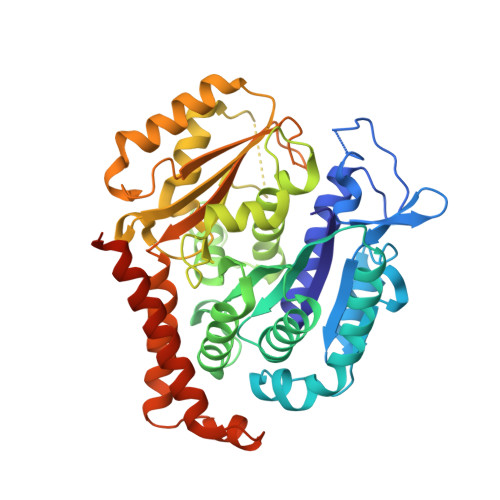
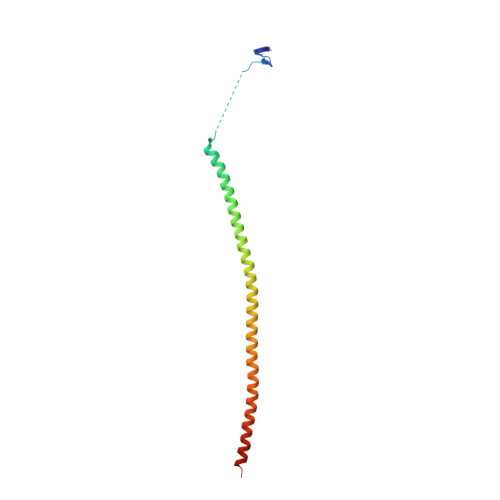
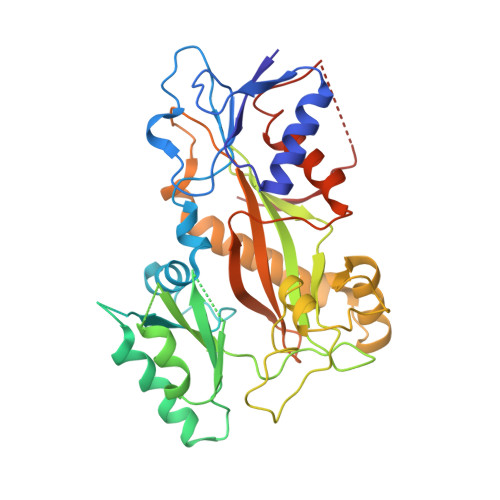
Microtubule (MT) dynamic behaviour is an attractive drug target for chemotherapy, whose regulation by MT-stabilizing and destabilizing agents has been fruitfully applied in treating several types of cancers. MT-stabilizing agents are also emerging as potential remedies for neurodegenerative conditions, such as Alzheimer's and Parkinson's disease, although single-target drugs are not expected to fully cure these complex pathologies. Drug combination often displays enhanced efficacy with respect to mono-therapies. In particular, MT-targeting bivalent compounds (MTBCs) represent a promising class of molecules; however, surprisingly, the majority of MTBCs reported so far exhibit equal if not less efficacy than their building monomers. In order to shed light on MTBCs poor performance, we characterised through a set of complementary approaches thiocolchine (TH) and two bivalent TH-homodimers as prototype molecules. First, the binding affinities of these three molecules were assessed, then we obtained the crystallographic structure of a tubulin-TH complex. The binding affinities were interpreted in light of structural data and of molecular dynamics simulations. Finally, their effects on MT cytoskeleton and cell survival were validated on HeLa cells. The ensemble of these data provides chemical and structural considerations on how a successful rational design of MTBCs should be conceived.
- Dipartimento di Bioscienze, Università degli Studi di Milano, via Celoria 26, 20133 Milano, Italy.
Organizational Affiliation: