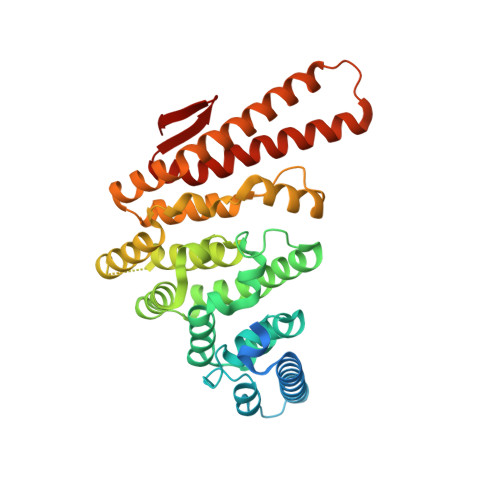
A unique surface on Pat1 C-terminal domain directly interacts with Dcp2 decapping enzyme and Xrn1 5'-3' mRNA exonuclease in yeast.
Charenton, C., Gaudon-Plesse, C., Fourati, Z., Taverniti, V., Back, R., Kolesnikova, O., Seraphin, B., Graille, M.(2017) Proc Natl Acad Sci U S A 114: E9493-E9501
- PubMed: 29078363 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1711680114
- Primary Citation Related Structures:
5LM5, 5LMF, 5LMG - PubMed Abstract:
The Pat1 protein is a central player of eukaryotic mRNA decay that has also been implicated in translational control. It is commonly considered a central platform responsible for the recruitment of several RNA decay factors. We demonstrate here that a yeast-specific C-terminal region from Pat1 interacts with several short motifs, named helical leucine-rich motifs (HLMs), spread in the long C-terminal region of yeast Dcp2 decapping enzyme. Structures of Pat1-HLM complexes reveal the basis for HLM recognition by Pat1. We also identify a HLM present in yeast Xrn1, the main 5'-3' exonuclease involved in mRNA decay. We show further that the ability of yeast Pat1 to bind HLMs is required for efficient growth and normal mRNA decay. Overall, our analyses indicate that yeast Pat1 uses a single binding surface to successively recruit several mRNA decay factors and show that interaction between those factors is highly polymorphic between species.
- Laboratoire de Biochimie, Ecole Polytechnique, CNRS, Université Paris-Saclay, 91128 Palaiseau cedex, France.
Organizational Affiliation: