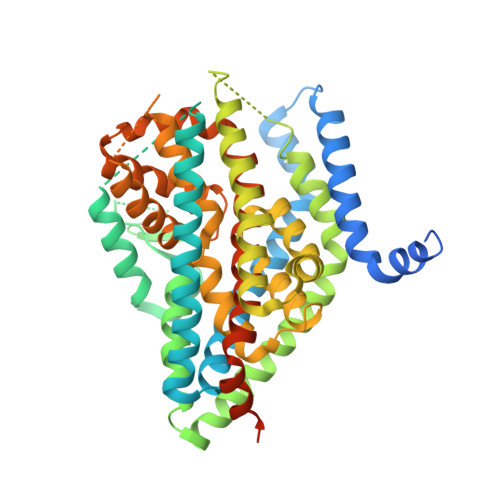
Structure and allosteric inhibition of excitatory amino acid transporter 1.
Canul-Tec, J.C., Assal, R., Cirri, E., Legrand, P., Brier, S., Chamot-Rooke, J., Reyes, N.(2017) Nature 544: 446-451
- PubMed: 28424515 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature22064
- Primary Citation Related Structures:
5LLM, 5LLU, 5LM4, 5MJU - PubMed Abstract:
Human members of the solute carrier 1 (SLC1) family of transporters take up excitatory neurotransmitters in the brain and amino acids in peripheral organs. Dysregulation of the function of SLC1 transporters is associated with neurodegenerative disorders and cancer. Here we present crystal structures of a thermostabilized human SLC1 transporter, the excitatory amino acid transporter 1 (EAAT1), with and without allosteric and competitive inhibitors bound. The structures reveal architectural features of the human transporters, such as intra- and extracellular domains that have potential roles in transport function, regulation by lipids and post-translational modifications. The coordination of the allosteric inhibitor in the structures and the change in the transporter dynamics measured by hydrogen-deuterium exchange mass spectrometry reveal a mechanism of inhibition, in which the transporter is locked in the outward-facing states of the transport cycle. Our results provide insights into the molecular mechanisms underlying the function and pharmacology of human SLC1 transporters.
- Molecular Mechanisms of Membrane Transport Laboratory, Institut Pasteur, 25-28 rue du Docteur Roux, 75015 Paris, France.
Organizational Affiliation: