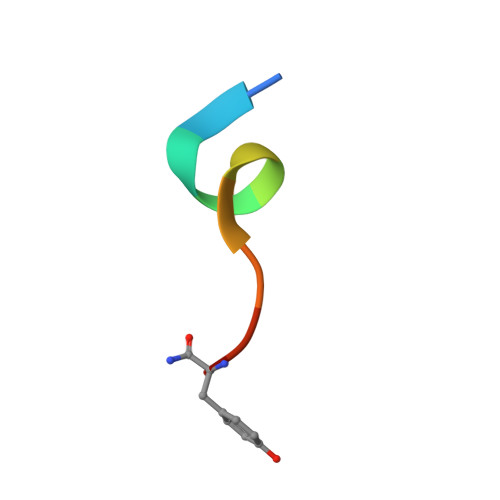
A Binuclear Zinc Interaction Fold Discovered in the Homodimer of Alzheimer's Amyloid-beta Fragment with Taiwanese Mutation D7H.
Polshakov, V.I., Mantsyzov, A.B., Kozin, S.A., Adzhubei, A.A., Zhokhov, S.S., van Beek, W., Kulikova, A.A., Indeykina, M.I., Mitkevich, V.A., Makarov, A.A.(2017) Angew Chem Int Ed Engl 56: 11734-11739
- PubMed: 28570778 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201704615
- Primary Citation Related Structures:
5LFY - PubMed Abstract:
Zinc-induced oligomerization of amyloid-β peptide (Aβ) produces potentially pathogenic agents of Alzheimer's disease. Mutations and modifications in the metal binding domain 1-16 of Aβ peptide crucially affect its zinc-induced oligomerization by changing intermolecular zinc mediated interface. The 3D structure of this interface appearing in a range of Aβ species is a prospective drug target for disease modifying therapy. Using NMR spectroscopy, EXAFS spectroscopy, mass spectrometry, and isothermal titration calorimetry the interaction of zinc ions with Aβ fragments 1-7 and 1-10 carrying familial Taiwanese mutation D7H was studied. Zinc ions induce formation of a stable homodimer formed by the two peptide chains fastened by two zinc ions and stacking interactions of imidazole rings. A binuclear zinc interaction fold in the dimer structure was discovered. It can be used for designing zinc-regulated proteins and zinc-mediated self-assembling peptides.
- Engelhardt Institute of Molecular Biology, Russian Academy of Sciences, 32 Vavilova str., Moscow, 119991, Russia.
Organizational Affiliation: