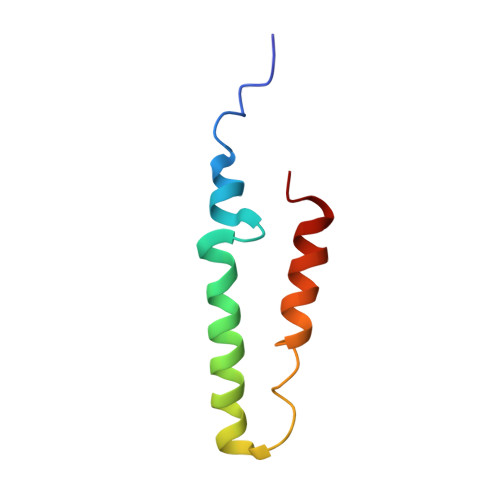
Structure of the Bacterial Sex F Pilus Reveals an Assembly of a Stoichiometric Protein-Phospholipid Complex.
Costa, T.R., Ilangovan, A., Ukleja, M., Redzej, A., Santini, J.M., Smith, T.K., Egelman, E.H., Waksman, G.(2016) Cell 166: 1436-1444.e10
- PubMed: 27610568 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2016.08.025
- Primary Citation Related Structures:
5LEG, 5LER, 5LFB - PubMed Abstract:
Conjugative pili are widespread bacterial appendages that play important roles in horizontal gene transfer, in spread of antibiotic resistance genes, and as sites of phage attachment. Among conjugative pili, the F "sex" pilus encoded by the F plasmid is the best functionally characterized, and it is also historically the most important, as the discovery of F-plasmid-mediated conjugation ushered in the era of molecular biology and genetics. Yet, its structure is unknown. Here, we present atomic models of two F family pili, the F and pED208 pili, generated from cryoelectron microscopy reconstructions at 5.0 and 3.6 Å resolution, respectively. These structures reveal that conjugative pili are assemblies of stoichiometric protein-phospholipid units. We further demonstrate that each pilus type binds preferentially to particular phospholipids. These structures provide the molecular basis for F pilus assembly and also shed light on the remarkable properties of conjugative pili in bacterial secretion and phage infection.
- Institute of Structural and Molecular Biology, University College London and Birkbeck, Malet Street, London WC1E 7HX, UK.
Organizational Affiliation: