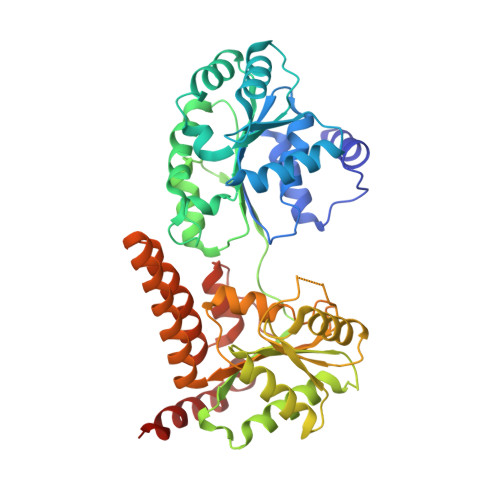
Crystal structure of human RECQL5 helicase in complex with 3D fragment (1-cyclohexyl-3-(oxolan-2-ylmethyl)urea)
Newman, J.A., Talon, R., Aitkenhead, H., Savitsky, P., Krojer, T., von Delft, F., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Gileadi, O.To be published.