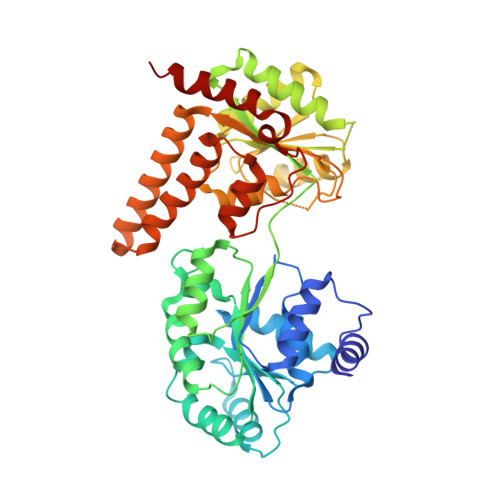
Insights into the RecQ helicase mechanism revealed by the structure of the helicase domain of human RECQL5.
Newman, J.A., Aitkenhead, H., Savitsky, P., Gileadi, O.(2017) Nucleic Acids Res 45: 4231-4243
- PubMed: 28100692 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkw1362
- Primary Citation Related Structures:
5LB3, 5LB5, 5LB8 - PubMed Abstract:
RecQ helicases are important maintainers of genome integrity with distinct roles in almost every cellular process requiring access to DNA. RECQL5 is one of five human RecQ proteins and is particularly versatile in this regard, forming protein complexes with a diverse set of cellular partners in order to coordinate its helicase activity to various processes including replication, recombination and DNA repair. In this study, we have determined crystal structures of the core helicase domain of RECQL5 both with and without the nucleotide ADP in two distinctly different ('Open' and 'Closed') conformations. Small angle X-ray scattering studies show that the 'Open' form of the protein predominates in solution and we discuss implications of this with regards to the RECQL5 mechanism and conformational changes. We have measured the ATPase, helicase and DNA binding properties of various RECQL5 constructs and variants and discuss the role of these regions and residues in the various RECQL5 activities. Finally, we have performed a systematic comparison of the RECQL5 structures with other RecQ family structures and based on these comparisons we have constructed a model for the mechano-chemical cycle of the common catalytic core of these helicases.
- Structural Genomics Consortium, University of Oxford, ORCRB, Roosevelt Drive, Oxford OX3 7DQ, UK.
Organizational Affiliation: