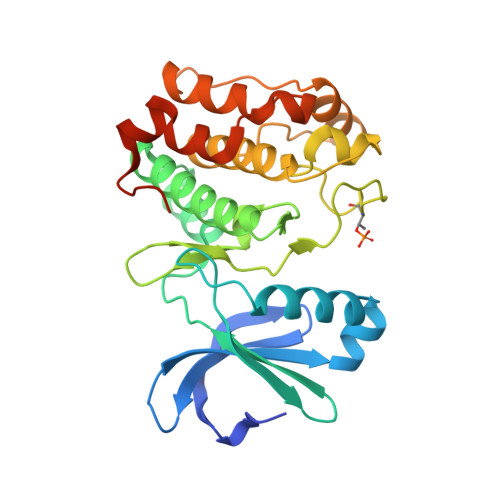
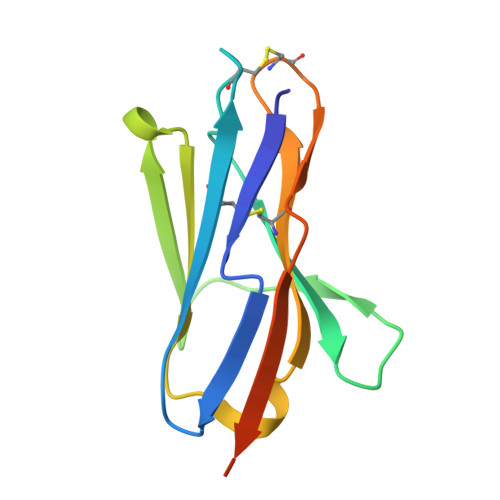
Allosteric inhibition of Aurora-A kinase by a synthetic vNAR domain.
Burgess, S.G., Oleksy, A., Cavazza, T., Richards, M.W., Vernos, I., Matthews, D., Bayliss, R.(2016) Open Biol 6
- PubMed: 27411893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1098/rsob.160089
- Primary Citation Related Structures:
5L8J, 5L8K, 5L8L - PubMed Abstract:
The vast majority of clinically approved protein kinase inhibitors target the ATP-binding pocket directly. Consequently, many inhibitors have broad selectivity profiles and most have significant off-target effects. Allosteric inhibitors are generally more selective, but are difficult to identify because allosteric binding sites are often unknown or poorly characterized. Aurora-A is activated through binding of TPX2 to an allosteric site on the kinase catalytic domain, and this knowledge could be exploited to generate an inhibitor. Here, we generated an allosteric inhibitor of Aurora-A kinase based on a synthetic, vNAR single domain scaffold, vNAR-D01. Biochemical studies and a crystal structure of the Aurora-A/vNAR-D01 complex show that the vNAR domain overlaps with the TPX2 binding site. In contrast with the binding of TPX2, which stabilizes an active conformation of the kinase, binding of the vNAR domain stabilizes an inactive conformation, in which the αC-helix is distorted, the canonical Lys-Glu salt bridge is broken and the regulatory (R-) spine is disrupted by an additional hydrophobic side chain from the activation loop. These studies illustrate how single domain antibodies can be used to characterize the regulatory mechanisms of kinases and provide a rational basis for structure-guided design of allosteric Aurora-A kinase inhibitors.
- Astbury Centre for Structural Molecular Biology, Faculty of Biological Sciences, University of Leeds, Leeds LS2 9JT, UK Department of Molecular and Cell Biology, University of Leicester, Leicester LE1 9HN, UK.
Organizational Affiliation: