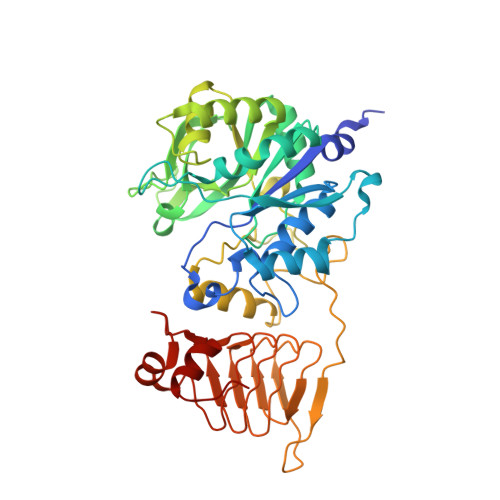
Structural Basis of Glycogen Biosynthesis Regulation in Bacteria.
Cifuente, J.O., Comino, N., Madariaga-Marcos, J., Lopez-Fernandez, S., Garcia-Alija, M., Agirre, J., Albesa-Jove, D., Guerin, M.E.(2016) Structure 24: 1613-1622
- PubMed: 27545622 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2016.06.023
- Primary Citation Related Structures:
5L6S, 5L6V - PubMed Abstract:
ADP-glucose pyrophosphorylase (AGPase) catalyzes the rate-limiting step of bacterial glycogen and plant starch biosynthesis, the most common carbon storage polysaccharides in nature. A major challenge is to understand how AGPase activity is regulated by metabolites in the energetic flux within the cell. Here we report crystal structures of the homotetrameric AGPase from Escherichia coli in complex with its physiological positive and negative allosteric regulators, fructose-1,6-bisphosphate (FBP) and AMP, and sucrose in the active site. FBP and AMP bind to partially overlapping sites located in a deep cleft between glycosyltransferase A-like and left-handed β helix domains of neighboring protomers, accounting for the fact that sensitivity to inhibition by AMP is modulated by the concentration of the activator FBP. We propose a model in which the energy reporters regulate EcAGPase catalytic activity by intra-protomer interactions and inter-protomer crosstalk, with a sensory motif and two regulatory loops playing a prominent role.
- Structural Biology Unit, CIC bioGUNE, Bizkaia Technology Park, 48160 Derio, Spain; Unidad de Biofísica, Centro Mixto Consejo Superior de Investigaciones Científicas, Universidad del País Vasco/Euskal Herriko Unibertsitatea (CSIC,UPV/EHU), Barrio Sarriena s/n, Leioa, Bizkaia, 48940, Spain; Departamento de Bioquímica, Universidad del País Vasco, 48940 Leioa, Bizkaia, Spain.
Organizational Affiliation: