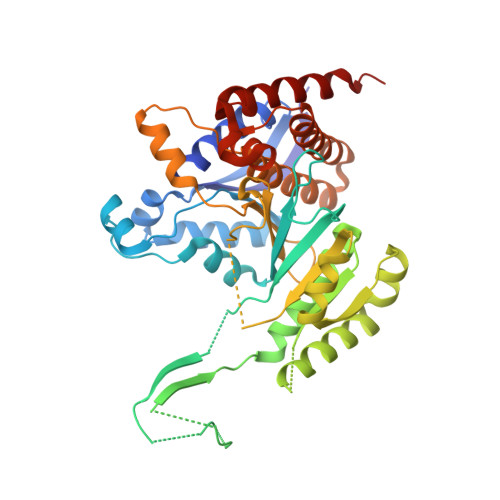
Discovery and Optimization of Allosteric Inhibitors of Mutant Isocitrate Dehydrogenase 1 (R132H IDH1) Displaying Activity in Human Acute Myeloid Leukemia Cells.
Jones, S., Ahmet, J., Ayton, K., Ball, M., Cockerill, M., Fairweather, E., Hamilton, N., Harper, P., Hitchin, J., Jordan, A., Levy, C., Lopez, R., McKenzie, E., Packer, M., Plant, D., Simpson, I., Simpson, P., Sinclair, I., Somervaille, T.C., Small, H., Spencer, G.J., Thomson, G., Tonge, M., Waddell, I., Walsh, J., Waszkowycz, B., Wigglesworth, M., Wiseman, D.H., Ogilvie, D.(2016) J Med Chem 59: 11120-11137
- PubMed: 28002956 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01320
- Primary Citation Related Structures:
5L57, 5L58 - PubMed Abstract:
A collaborative high throughput screen of 1.35 million compounds against mutant (R132H) isocitrate dehydrogenase IDH1 led to the identification of a novel series of inhibitors. Elucidation of the bound ligand crystal structure showed that the inhibitors exhibited a novel binding mode in a previously identified allosteric site of IDH1 (R132H). This information guided the optimization of the series yielding submicromolar enzyme inhibitors with promising cellular activity. Encouragingly, one compound from this series was found to induce myeloid differentiation in primary human IDH1 R132H AML cells in vitro.
- Manchester Institute of Biotechnology, University of Manchester , Princess Street, Manchester, M1 7DN, U.K.
Organizational Affiliation: