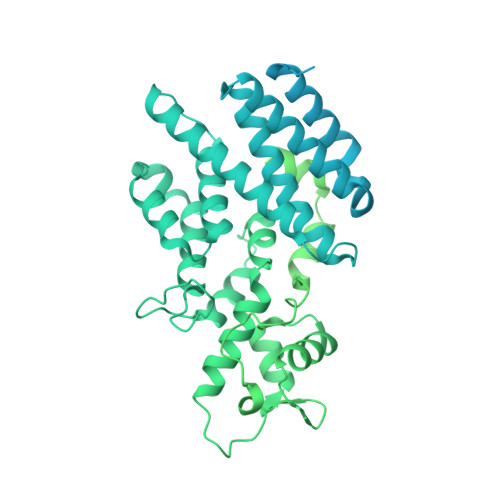
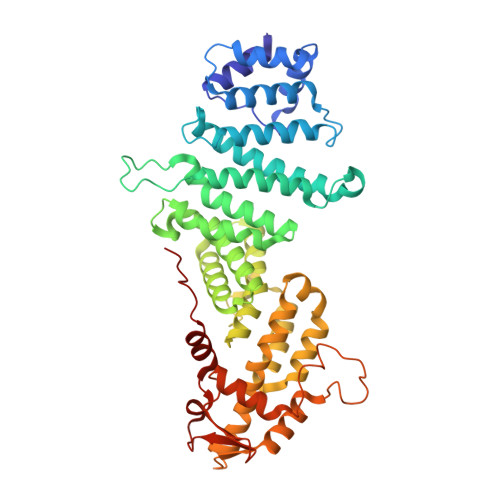
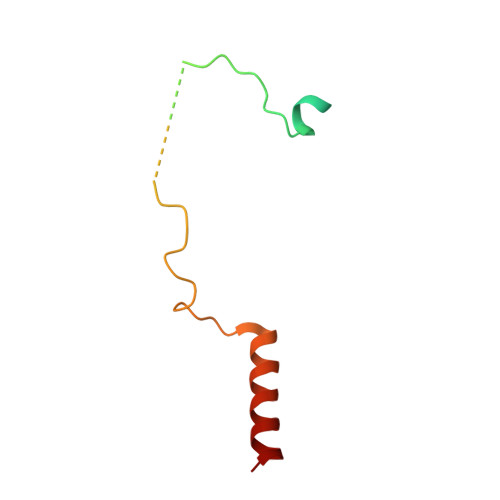
The Sac3 TPR-like region in the Saccharomyces cerevisiae TREX-2 complex is more extensive but independent of the CID region.
Aibara, S., Bai, X.C., Stewart, M.(2016) J Struct Biol 195: 316-324
- PubMed: 27422657
- DOI: https://doi.org/10.1016/j.jsb.2016.07.007
- Primary Citation Related Structures:
5G5P, 5L3T - PubMed Abstract:
Transcription-export complex 2 (TREX-2 complex) facilitates the localization of actively transcribing genes to the nuclear periphery and also functions to contribute to the generation of export-competent mRNPs through interactions with the general mRNA nuclear export factor Mex67:Mtr2. The TREX-2 complex is based on a Sac3 scaffold to which Thp1, Sem1, Cdc31, and Sus1 bind. TREX-2 can be subdivided into two modules: one, in which Thp1 and Sem1 bind to the Sac3(M) region (residues ∼100-551), and the other in which Cdc31 and two Sus1 chains bind to the Sac3(CID) region (residues ∼710-805). Complementary structural analyses using X-ray crystallography, electron microscopy, and small-angle X-ray scattering of the Saccharomyces cerevisiae TREX-2 complex, expressed using Baculovirus-infected Sf9 cells, have indicated that the TPR-like repeats of the Sac3(M) region extend considerably further towards the N-terminus than previously thought, and also indicate that this region and Sac3(CID):Sus1:Cdc31 region of the S. cerevisiae complex are structurally independent. Although the density visible accounted for only ∼100kDa, a 5.3Å resolution cryo-EM reconstruction was obtained of the M-region of TREX-2 that showed an additional three putative α-helices extending towards the Sac3 N-terminus and these helices were also seen in a 4.9Å resolution structure obtained by X-ray crystallography. We describe the expression, purification and structural characterization of the S. cerevisiae TREX-2 complex and demonstrate that the Sac3 TPR-like repeats are more extensive than previously thought and that the M- and CID-regions do not appear to have a defined spatial orientation.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge Biomedical Campus, Cambridge CB2 0QH, United Kingdom.
Organizational Affiliation: