Discovery of Benzothiazole Scaffold-Based DNA Gyrase B Inhibitors.
Gjorgjieva, M., Tomasic, T., Barancokova, M., Katsamakas, S., Ilas, J., Tammela, P., Peterlin Masic, L., Kikelj, D.(2016) J Med Chem 59: 8941-8954
- PubMed: 27541007 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00864
- Primary Citation Related Structures:
5L3J - PubMed Abstract:
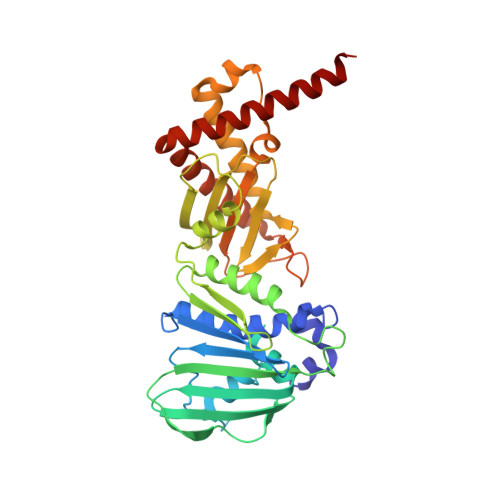
Bacterial DNA gyrase and topoisomerase IV control the topological state of DNA during replication and are validated targets for antibacterial drug discovery. Starting from our recently reported 4,5,6,7-tetrahydrobenzo[1,2-d]thiazole-based DNA gyrase B inhibitors, we replaced their central core with benzothiazole-2,6-diamine scaffold and interchanged substituents in positions 2 and 6. This resulted in equipotent nanomolar inhibitors of DNA gyrase from Escherichia coli displaying improved inhibition of Staphylococcus aureus DNA gyrase and topoisomerase IV from both bacteria. Compound 27 was the most balanced inhibitor of DNA gyrase and topoisomerase IV from both E. coli and S. aureus. The crystal structure of the 2-((2-(4,5-dibromo-1H-pyrrole-2-carboxamido)benzothiazol-6-yl)amino)-2-oxoacetic acid (24) in complex with E. coli DNA gyrase B revealed the binding mode of the inhibitor in the ATP-binding pocket. Only some compounds possessed weak antibacterial activity against Gram-positive bacteria. These results provide a basis for structure-based optimization toward dual DNA gyrase and topoisomerase IV inhibitors with antibacterial activity.
- Faculty of Pharmacy, University of Ljubljana , Aškerčeva 7, 1000 Ljubljana, Slovenia.
Organizational Affiliation: