Novel Autotaxin Inhibitors for the Treatment of Osteoarthritis Pain: Lead Optimization via Structure-Based Drug Design.
Jones, S.B., Pfeifer, L.A., Bleisch, T.J., Beauchamp, T.J., Durbin, J.D., Klimkowski, V.J., Hughes, N.E., Rito, C.J., Dao, Y., Gruber, J.M., Bui, H., Chambers, M.G., Chandrasekhar, S., Lin, C., McCann, D.J., Mudra, D.R., Oskins, J.L., Swearingen, C.A., Thirunavukkarasu, K., Norman, B.H.(2016) ACS Med Chem Lett 7: 857-861
- PubMed: 27660691 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.6b00207
- Primary Citation Related Structures:
5L0B, 5L0E, 5L0K - PubMed Abstract:
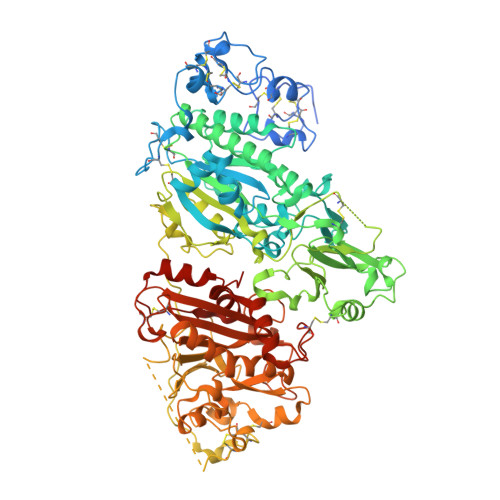
In an effort to develop a novel therapeutic agent aimed at addressing the unmet need of patients with osteoarthritis pain, we set out to develop an inhibitor for autotaxin with excellent potency and physical properties to allow for the clinical investigation of autotaxin-induced nociceptive and neuropathic pain. An initial hit identification campaign led to an aminopyrimidine series with an autotaxin IC50 of 500 nM. X-ray crystallography enabled the optimization to a lead compound that demonstrated favorable potency (IC50 = 2 nM), PK properties, and a robust PK/PD relationship.
- Lilly Research Laboratories , A Division of Eli Lilly and Company, Indianapolis, Indiana 46285, United States.
Organizational Affiliation: