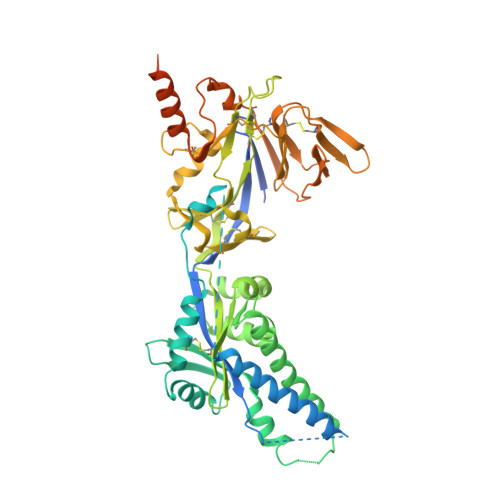
Therapeutic efficacy of a respiratory syncytial virus fusion inhibitor.
Roymans, D., Alnajjar, S.S., Battles, M.B., Sitthicharoenchai, P., Furmanova-Hollenstein, P., Rigaux, P., Berg, J.V.D., Kwanten, L., Ginderen, M.V., Verheyen, N., Vranckx, L., Jaensch, S., Arnoult, E., Voorzaat, R., Gallup, J.M., Larios-Mora, A., Crabbe, M., Huntjens, D., Raboisson, P., Langedijk, J.P., Ackermann, M.R., McLellan, J.S., Vendeville, S., Koul, A.(2017) Nat Commun 8: 167-167
- PubMed: 28761099 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-00170-x
- Primary Citation Related Structures:
5KWW - PubMed Abstract:
Respiratory syncytial virus is a major cause of acute lower respiratory tract infection in young children, immunocompromised adults, and the elderly. Intervention with small-molecule antivirals specific for respiratory syncytial virus presents an important therapeutic opportunity, but no such compounds are approved today. Here we report the structure of JNJ-53718678 bound to respiratory syncytial virus fusion (F) protein in its prefusion conformation, and we show that the potent nanomolar activity of JNJ-53718678, as well as the preliminary structure-activity relationship and the pharmaceutical optimization strategy of the series, are consistent with the binding mode of JNJ-53718678 and other respiratory syncytial virus fusion inhibitors. Oral treatment of neonatal lambs with JNJ-53718678, or with an equally active close analog, efficiently inhibits established acute lower respiratory tract infection in the animals, even when treatment is delayed until external signs of respiratory syncytial virus illness have become visible. Together, these data suggest that JNJ-53718678 is a promising candidate for further development as a potential therapeutic in patients at risk to develop respiratory syncytial virus acute lower respiratory tract infection.Respiratory syncytial virus causes lung infections in children, immunocompromised adults, and in the elderly. Here the authors show that a chemical inhibitor to a viral fusion protein is effective in reducing viral titre and ameliorating infection in rodents and neonatal lambs.
- Janssen Infectious Diseases and Vaccines, Janssen Pharmaceutica NV, Turnhoutseweg 30, 2340, Beerse, Belgium. droymans@its.jnj.com.
Organizational Affiliation: