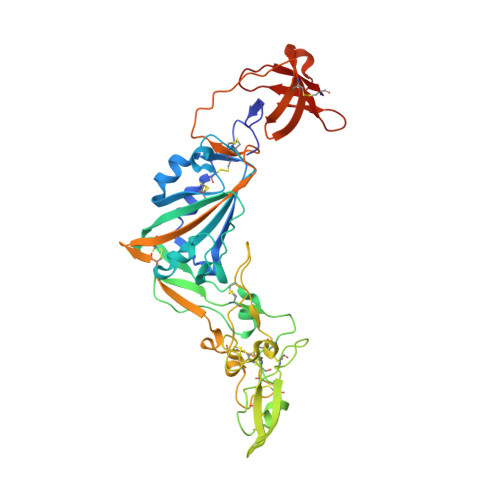
Crystal structure of the receptor binding domain of the spike glycoprotein of human betacoronavirus HKU1
Ou, X., Guan, H., Qin, B., Mu, Z., Wojdyla, J.A., Wang, M., Dominguez, S.R., Qian, Z., Cui, S.(2017) Nat Commun 8: 15216-15216
- PubMed: 28534504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15216
- Primary Citation Related Structures:
5GNB, 5KWB - PubMed Abstract:
Human coronavirus (CoV) HKU1 is a pathogen causing acute respiratory illnesses and so far little is known about its biology. HKU1 virus uses its S1 subunit C-terminal domain (CTD) and not the N-terminal domain like other lineage A β-CoVs to bind to its yet unknown human receptor. Here we present the crystal structure of HKU1 CTD at 1.9 Å resolution. The structure consists of three subdomains: core, insertion and subdomain-1 (SD-1). While the structure of the core and SD-1 subdomains of HKU1 are highly similar to those of other β-CoVs, the insertion subdomain adopts a novel fold, which is largely invisible in the cryo-EM structure of the HKU1 S trimer. We identify five residues in the insertion subdomain that are critical for binding of neutralizing antibodies and two residues essential for receptor binding. Our study contributes to a better understanding of entry, immunity and evolution of CoV S proteins.
- MOH Key Laboratory of Systems Biology of Pathogens, Institute of Pathogen Biology, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing 100730, China.
Organizational Affiliation: