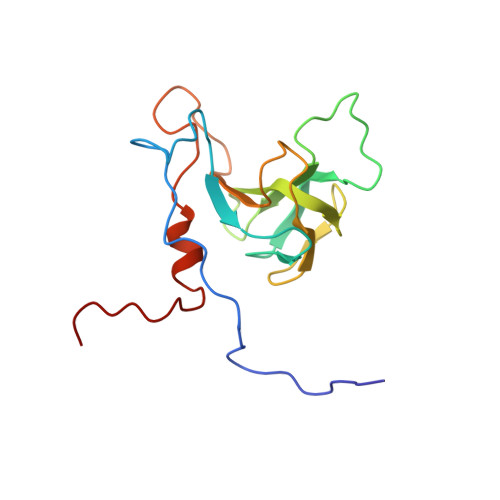
Solution structure of the Cys74 to Ala74 mutant of the recombinant catalytic domain of Zoocin A.
Xing, M., Simmonds, R.S., Timkovich, R.(2017) Proteins 85: 177-181
- PubMed: 27699884 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25178
- Primary Citation Related Structures:
5KVP - PubMed Abstract:
Zoocin A is a Zn-metallopeptidase secreted by Streptococcus zooepidemicus strain 4881. Its catalytic domain is responsible for cleaving the D-alanyl-L-alanine peptide bond in streptococcal peptidoglycan. The solution NMR structure of the Cys74 to Ala74 mutant of the recombinant catalytic domain (rCAT C74A) has been determined. With a previous structure determination for the recombinant target recognition domain (rTRD), this completes the 3D structure of zoocin A. While the structure of rCAT C74A resembles those of the catalytic domains of lysostaphin and LytM, the substrate binding groove is wider and no tyrosine residue was observed in the active site. Proteins 2016; 85:177-181. © 2016 Wiley Periodicals, Inc.
- Department of Chemistry, University of Alabama, Tuscaloosa, Alabama, 35487.
Organizational Affiliation: